Now Reading: Bacteria That Breathe Rocks And Sulfur
-
01
Bacteria That Breathe Rocks And Sulfur
Bacteria That Breathe Rocks And Sulfur
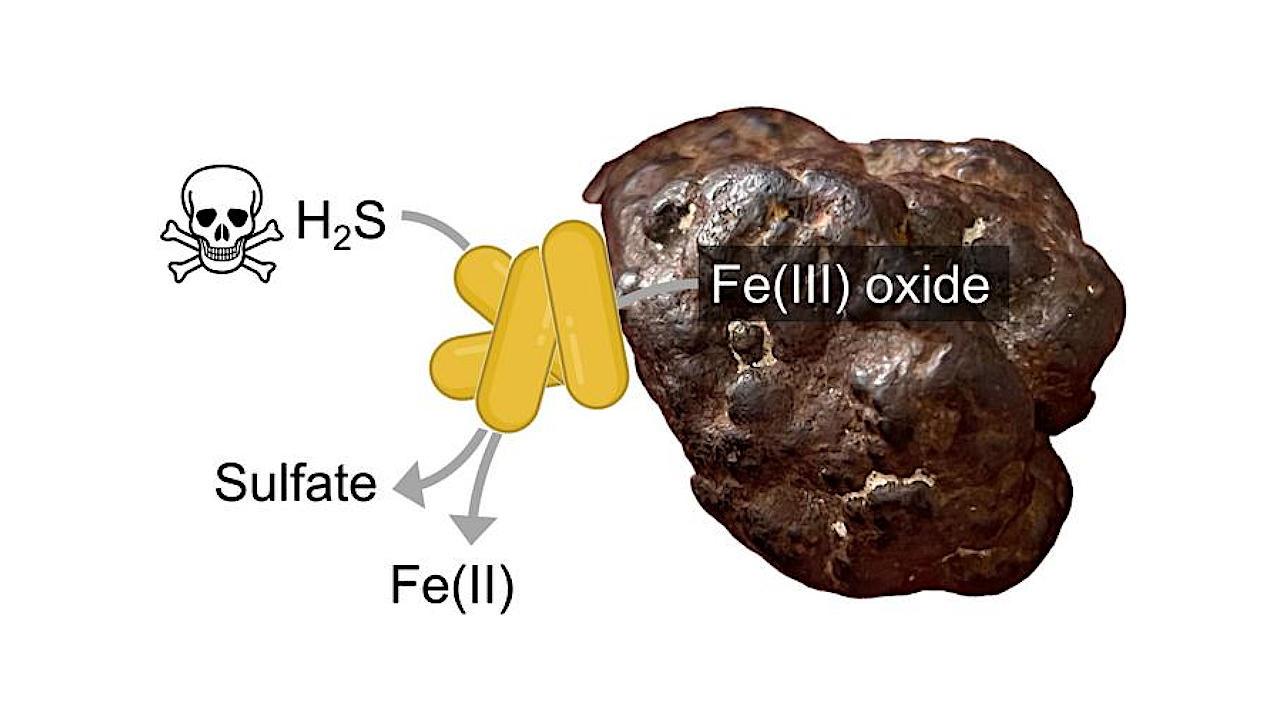

Scheme of MISO bacteria on an iron(III) oxide mineral particle. Credit Alexander Loy
An international team of scientists led by microbiologists Marc Mussmann and Alexander Loy from the University of Vienna has discovered a new microbial metabolism: so-called MISO bacteria “breathe” iron minerals by oxidizing toxic sulfide.
The researchers found that the reaction between toxic hydrogen sulfide and solid iron minerals is not only a chemical process, but also a previously unknown biological process in which versatile microbes in marine sediments and terrestrial wetlands remove toxic sulfide and use it for their growth. These bacteria could prevent the spread of oxygen-free “dead zones” in aquatic environments. The findings have now been published in Nature.
The global element cycles
The biogeochemical cycles of carbon, nitrogen, sulfur, and iron describe how these elements are transformed through reduction and oxidation (redox) reactions and how they move between the atmosphere, water, soil, rocks, and living organisms. These cycles are intricately linked to Earth’s climate, as they regulate the flow of greenhouse gases and influence the planet’s temperature balance. Microorganisms are central to nearly every step of these redox transformations, using compounds like sulfur and iron for respiration in much the same way humans use oxygen to metabolize food.
Sulfur and iron are particularly vital for microbial life in oxygen-deprived environments, such as the ocean floor or wetlands. Sulfur exists in various forms — as a gas in the atmosphere, as sulfate in oceans, or as part of minerals in rocks. Similarly, iron can transition between different forms depending on the presence of oxygen. When microbes metabolize sulfur, they often alter the form of iron simultaneously, and vice versa. This coupling of sulfur and iron cycles has far-reaching implications, influencing nutrient availability and the production or breakdown of greenhouse gases like carbon dioxide and methane. Understanding these interconnected cycles is crucial for predicting how ecosystems respond to pollution, climate change, and other human activities.
Breathing iron minerals to detoxify Sulfide
The activities of specialized microbes in oxygen-free environments such as marine sediments, wetlands, and freshwater aquifers produce hydrogen sulfide – a toxic gas with a distinctive rotten egg smell. The interaction between sulfide and solid iron(III) oxide minerals such as rusty iron plays a key role in controlling sulfide concentrations. Until now, biogeochemical models have treated this reaction as purely abiotic, primarily resulting in the formation of elemental sulfur and iron monosulfide (FeS), a black mineral that is, for example, responsible for dark coloration of beach sediments under low oxygen.
“We show that this environmentally important redox reaction is not solely chemical,” explains Alexander Loy, research group leader at CeMESS, the Centre for Microbiology and Environmental Systems Science at the University of Vienna. “Microorganisms can also harness it for growth.”
The newly discovered microbial energy metabolism, shortly termed MISO, couples the reduction of iron(III) oxide with the oxidation of sulfide. Unlike the chemical reaction, MISO directly produces sulfate, effectively bypassing intermediate steps in the sulfur cycle. “MISO bacteria remove toxic sulfide and may help prevent the expansion of so-called “dead zones” in aquatic environments, while fixing carbon dioxide for growth – similar to plants,” adds Marc Mussmann, senior scientist at CeMESS.
A globally important microbial process that outpaces chemistry
In laboratory growth experiments with a cultivated MISO bacterium, the researchers demonstrated that the enzymatically catalyzed reaction is faster than the equivalent chemical reaction. This suggests that microbes are the primary drivers of this process in nature. “Diverse bacteria and archaea possess the genetic capacity for MISO,” explains Song-Can Chen, lead author of the study, “and they are found in a wide range of natural and human-made environments.” In marine sediments, MISO could account for up to 7% of global sulfide oxidation to sulfate, driven by the substantial flux of reactive iron from rivers and melting glaciers into the oceans.
The findings of the University of Vienna team, which is supported by the Austrian Science Fund (FWF) as part of the ‘Microbiomes Drive Planetary Health’ Cluster of Excellence, reveal a previously unknown biological mechanism that links sulfur, iron, and carbon cycling in oxygen-free environments. “This discovery demonstrates the metabolic ingenuity of microorganisms and highlights their indispensable role in shaping Earth’s global element cycles.” Alexander Loy concludes.

D. alkaliphilus uses a reversed canonical dissimilatory sulfate reduction (Dsr) pathway to oxidize sulfide to sulfate (blue protein labels). Sulfide reacts with DsrC to form DsrC trisulfide (DCT), which is oxidized stepwise to sulfate via DsrAB, AprAB and Sat. The reducing equivalents are transferred to iron(iii) oxides via EET mechanisms (green protein labels) facilitated by MHCs. D. alkaliphilus also has genes encoding type IV pili (grey protein labels). The tight adhesion (Tad) pilus may drive the cell adherence to the surface of solid iron oxides and/or FeS. The transcription level of each gene during MISO is indicated by mean reads per kilobase of transcript per million mapped reads (RPKM) of four replicates corresponding to the colour in the legend. Blue and green dots show significantly increased gene transcription during sulfide/FeS oxidation and ferrihydrite reduction, respectively. The identifier, full name, RPKM, statistical significance of differential transcription and genomic arrangement for each gene are provided in Extended Data Fig. 3 and Supplementary Tables 2–5. Electron flow and proton translocation are indicated by red arrows. Black dotted arrows indicate putative sulfur redox reactions with unclear enzymology. APS, adenosine-5′-phosphosulfate, Qox, oxidized menaquinone; Qred, reduced menaquinone; FoxFeR, incubation of D. alkaliphilus under formate-oxidizing and ferrihydrite-reducing conditions; FeSoxFeR, incubation of D. alkaliphilus under FeS-oxidizing and ferrihydrite-reducing conditions; Sdisp, incubation of D. alkaliphilus under S(0) disproportionation conditions; SoxFeR, incubation of D. alkaliphilus under sulfide-oxidizing and ferrihydrite-reducing conditions; SoxNR, incubation of D. alkaliphilus under sulfide-oxidizing and nitrate-reducing conditions. — Nature
Microbial iron oxide respiration coupled to sulfide oxidation, Nature (open access)
Astrobiology
Stay Informed With the Latest & Most Important News
Previous Post
Next Post
-
 01Two Black Holes Observed Circling Each Other for the First Time
01Two Black Holes Observed Circling Each Other for the First Time -
 02From Polymerization-Enabled Folding and Assembly to Chemical Evolution: Key Processes for Emergence of Functional Polymers in the Origin of Life
02From Polymerization-Enabled Folding and Assembly to Chemical Evolution: Key Processes for Emergence of Functional Polymers in the Origin of Life -
 03Astronomy 101: From the Sun and Moon to Wormholes and Warp Drive, Key Theories, Discoveries, and Facts about the Universe (The Adams 101 Series)
03Astronomy 101: From the Sun and Moon to Wormholes and Warp Drive, Key Theories, Discoveries, and Facts about the Universe (The Adams 101 Series) -
 04True Anomaly hires former York Space executive as chief operating officer
04True Anomaly hires former York Space executive as chief operating officer -
 05Φsat-2 begins science phase for AI Earth images
05Φsat-2 begins science phase for AI Earth images -
 06Hurricane forecasters are losing 3 key satellites ahead of peak storm season − a meteorologist explains why it matters
06Hurricane forecasters are losing 3 key satellites ahead of peak storm season − a meteorologist explains why it matters -
 07Binary star systems are complex astronomical objects − a new AI approach could pin down their properties quickly
07Binary star systems are complex astronomical objects − a new AI approach could pin down their properties quickly