Now Reading: Minimal Amino Acid Alphabet for Protein Design
-
01
Minimal Amino Acid Alphabet for Protein Design
Minimal Amino Acid Alphabet for Protein Design
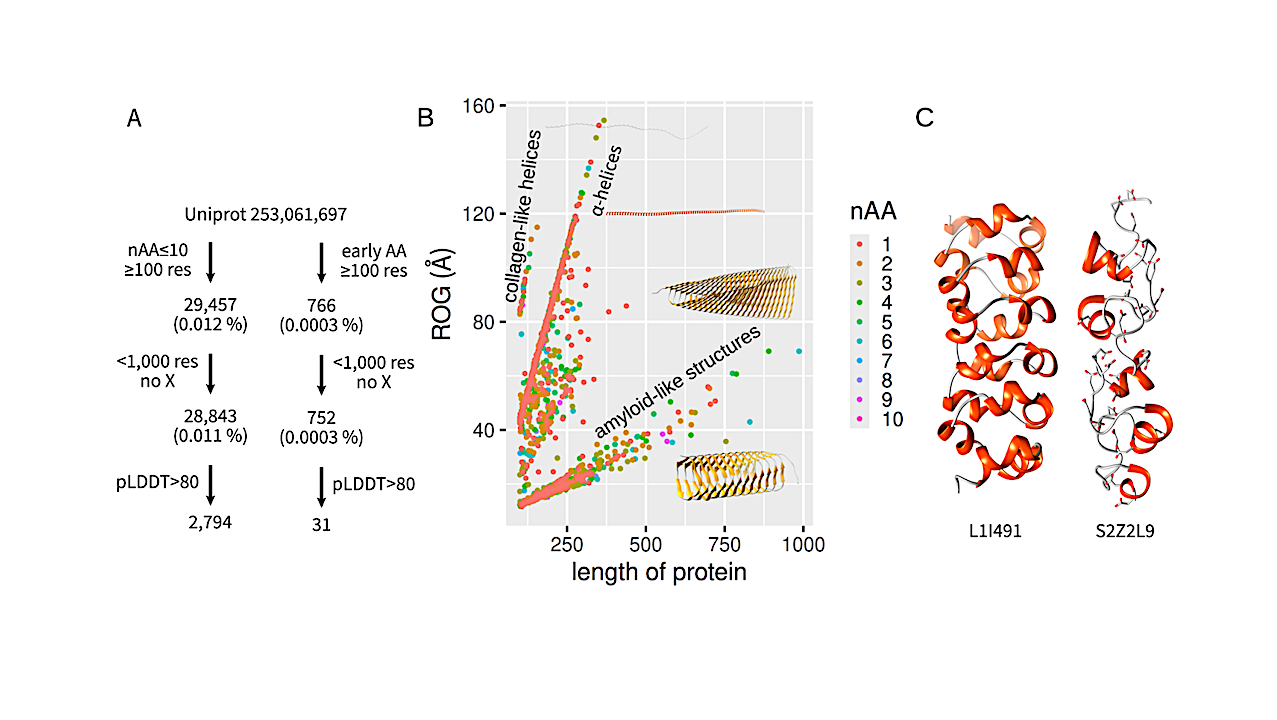

Utilization of reduced amino acid alphabets by modern proteins. (A) Workflow for finding of modern proteins composed of any reduced amino acid alphabet (left) or a reduced alphabet from early amino acids (right). (B) Analysis of the resulting 2,794 proteins. Each protein is represented as a dot colored by the number of amino acids from which it is formed. Lines pointing from the origin correspond to repetitive structures. (C) Selected interesting proteins L1I491 (uncharacterized protein from Guillardia theta) and S2Z2L9 thrombospondin type 3 repeat-containing protein from Corynebacterium pyruviciproducens. — biorxiv.org
Proteins are built from 20 canonical amino acids. It is interesting to explore whether proteins can be formed from significantly reduced amino acid alphabets.
Our bioinformatics survey of UniProt (more than 250 M sequences) revealed that proteins composed of reduced amino acid alphabets (< 10) are extremely rare among existing proteins.
Next, we used computational protein design to design proteins composed of all 1,013 possible alphabets of 2-10 early amino acids (Ala, Asp, Glu, Gly, Ile, Leu, Pro, Ser, Thr, and Val). The length of all proteins was 100 amino acid residues. Small amino acid alphabets preferred simple helices or helix bundles.
Larger amino acid alphabets allowed for the design of more complex structures. A protein composed of 8 amino acids (Ala, Asp, Gly, Leu, Val, Ser, Thr, and Pro) was successfully experimentally verified. It belongs to fibronectin type III domain β-sheet-rich architecture.
Attempts to experimentally verify designs composed of 6 and 4 amino acids were unsuccessful. We show by a computational experiment without an experimental validation that inverse folding programs, namely ProteinMPNN, can stabilize designed proteins within the same amino acid alphabet.
Our results show that globular proteins may have formed early in evolution. Furthermore, we show that it is possible to design proteins with interesting properties for biotechnology and synthetic biology.
Minimal Amino Acid Alphabet for Protein Design, biorxiv.org (open access)
Astrobiology, Genomics,
Stay Informed With the Latest & Most Important News
Previous Post
Next Post
-
 01Two Black Holes Observed Circling Each Other for the First Time
01Two Black Holes Observed Circling Each Other for the First Time -
 02From Polymerization-Enabled Folding and Assembly to Chemical Evolution: Key Processes for Emergence of Functional Polymers in the Origin of Life
02From Polymerization-Enabled Folding and Assembly to Chemical Evolution: Key Processes for Emergence of Functional Polymers in the Origin of Life -
 03Astronomy 101: From the Sun and Moon to Wormholes and Warp Drive, Key Theories, Discoveries, and Facts about the Universe (The Adams 101 Series)
03Astronomy 101: From the Sun and Moon to Wormholes and Warp Drive, Key Theories, Discoveries, and Facts about the Universe (The Adams 101 Series) -
 04True Anomaly hires former York Space executive as chief operating officer
04True Anomaly hires former York Space executive as chief operating officer -
 05Φsat-2 begins science phase for AI Earth images
05Φsat-2 begins science phase for AI Earth images -
 06Hurricane forecasters are losing 3 key satellites ahead of peak storm season − a meteorologist explains why it matters
06Hurricane forecasters are losing 3 key satellites ahead of peak storm season − a meteorologist explains why it matters -
 07Binary star systems are complex astronomical objects − a new AI approach could pin down their properties quickly
07Binary star systems are complex astronomical objects − a new AI approach could pin down their properties quickly