Now Reading: Anoxia-adapted Cyanobacteria In A Marine Blue Hole
-
01
Anoxia-adapted Cyanobacteria In A Marine Blue Hole
Anoxia-adapted Cyanobacteria In A Marine Blue Hole
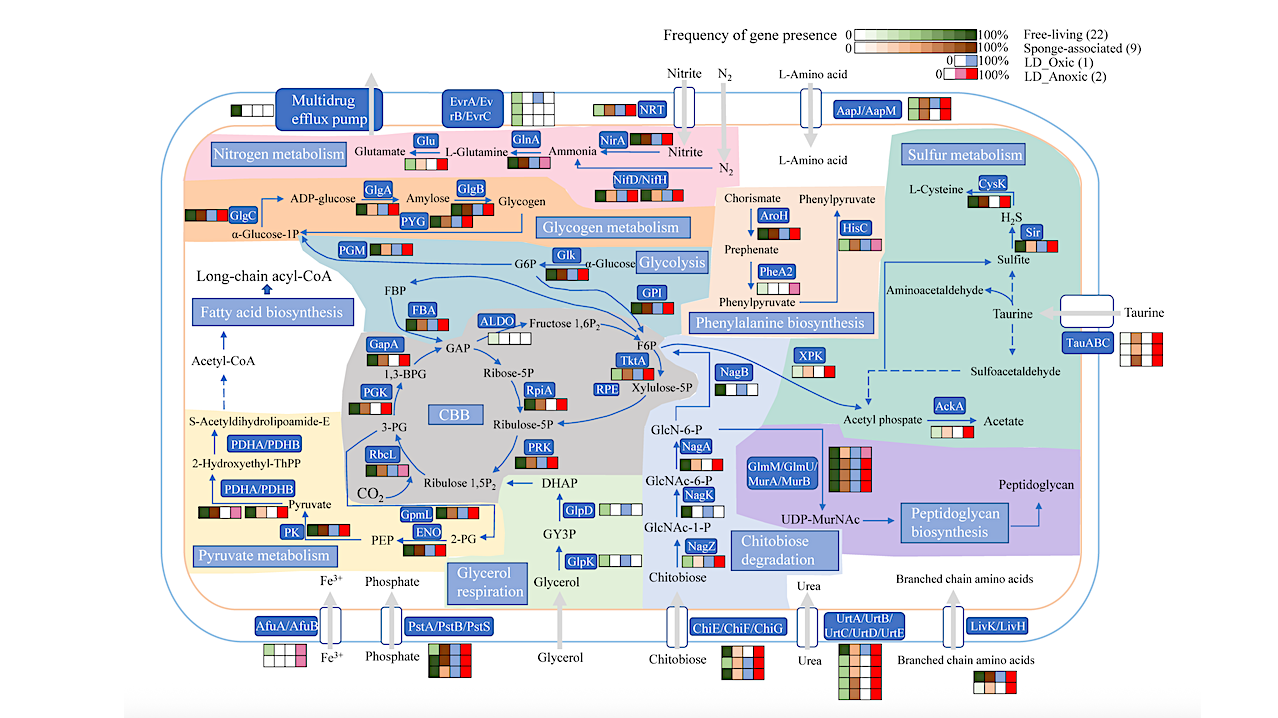

Metabolic reconstruction of major pathways in Synechococcus MAGs. The draft genomes for free-living and sponge-associated Synechococcus from NCBI were compared with the two MAGs from the anoxic YBH layer. The number or percentage of KEGG genes in these genomes was illustrated to mediate the metabolic network. α-Glucose-1P: α-Glucose-1-phosphate; G6P: α-D-Glucose 6-phosphate; F6P: D-Fructose 6-phosphate; FBP: D-Fructose 1,6-bisphosphate; 3-PG: 3-Phospho-D-glycerate; 1,3-BPG: 1,3-Bisphospho-D-glycerate; GAP: Glyceraldehyde 3-phosphate; Ribose-5P: Ribose 5-phosphate; Ribulose-5P: D-Ribulose 5-phosphate; Ribulose 1,5P2: D-Ribulose 1,5-bisphosphate; Fructose 1,6P2: D-Fructose 1,6-bisphosphate; Xylulose-5P: D-Xylulose 5-phosphate; PEP: Phosphoenolpyruvate; 2-PG: 2-Phospho-D-glycerate; DHAP: dihydroxyacetone phosphate. GY3P: glycerol-3-phosphate; GlcNAc-1-P: N-Acetyl-alpha-D-glucosamine 1-phosphate; GlcNAc-6-P: N-Acetyl-D-glucosamine 6-phosphate; GlcN-6-P: D-Glucosamine 6-phosphate. — Applied and Environmental Microbiology
Vertical transmission of marine particles brings ocean surface cyanobacteria into the deep ocean, where heterotrophic cyanobacterial lineages probably evolve to adapt to new environments even in oxygen-depleted zones.
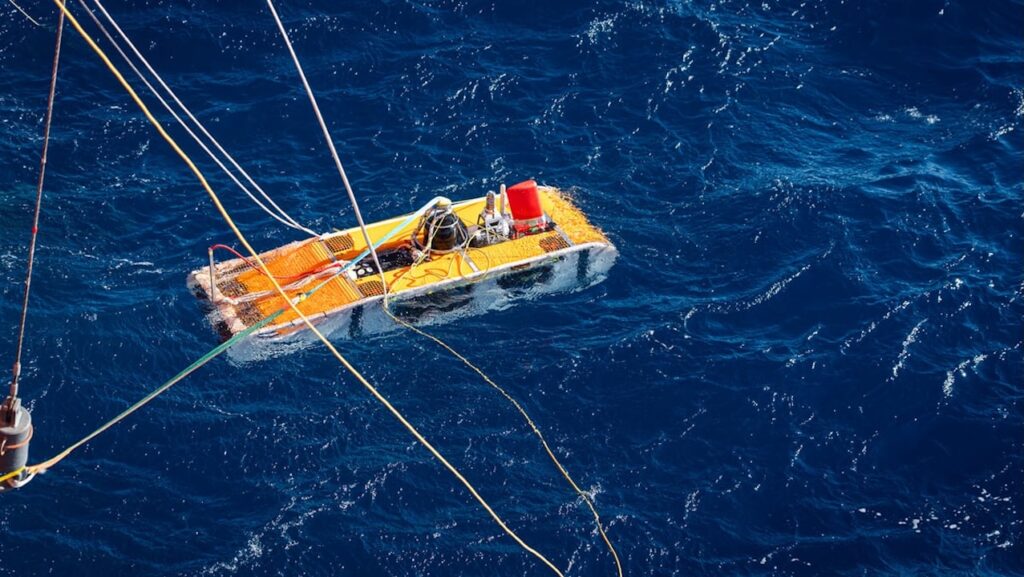
At present, active cyanobacteria have rarely been reported in dark and anoxic water columns in the deep sea. In this study, we recovered three metagenome-assembled genomes of cyanobacteria from the Yongle blue hole located in the South China Sea, two of which were actively transcribed in a dark, anoxic environment at 250 m depth, through integrated metagenomic and metatranscriptomic analyses of water samples from 21 stratified depths collected using in situ microbial fixation and filtration.

Relative abundance of Synechococcus MAGs in metagenomes and metatranscriptomes. (a) The division of oxic, suboxic, and anoxic zones in YBH. (b) The relative abundance of three Synechococcus MAGs across 21 metagenomes. (c) The relative abundance of three Synechococcus MAGs across 11 metatranscriptomes. Oxic-bin111 represents a Synechococcus genome reconstructed from the oxic layer, while Anoxic-bin48 and Anoxic-bin49 are two closely related Synechococcus genomes recovered from the anoxic layer. The samples marked with an asterisk (*) at the depth were collected in September, while the rest were collected in April 2021. — Applied and Environmental Microbiology
These anoxia-adapted cyanobacteria were phylogenetically approximate to the sponge cyanobacterial symbionts, while the genomic features showed similarities with both free-living and sponge symbiotic counterparts. They exhibit genomic features shared with symbiotic lineages, including loss of substrate utilization, biosynthesis pathways, DNA repair, and circadian regulation.
Conversely, they retain selected metabolic characteristics of free-living lineages, including phenylalanine biosynthesis and phosphoserine metabolism. Additionally, the discovery of taurine transport proteins in the genomes suggests the potential for organic sulfur uptake from the environment.
Altogether, these findings reveal a distinct genomic configuration in cyanobacteria inhabiting a permanently dark and anoxic marine system, characterized by the retention of oxygen-dependent metabolic potential alongside sustained transcriptional suppression under in situ conditions. This study provides new insights into the ecological persistence and evolutionary adaptation of cyanobacteria under long-term oxygen limitation.
Anoxia-adapted cyanobacteria in a marine blue hole, American Society for Microbiology
Astrobiology, genomics, extremophile,
Stay Informed With the Latest & Most Important News
Previous Post
Next Post
-
 01Two Black Holes Observed Circling Each Other for the First Time
01Two Black Holes Observed Circling Each Other for the First Time -
 02From Polymerization-Enabled Folding and Assembly to Chemical Evolution: Key Processes for Emergence of Functional Polymers in the Origin of Life
02From Polymerization-Enabled Folding and Assembly to Chemical Evolution: Key Processes for Emergence of Functional Polymers in the Origin of Life -
 03Astronomy 101: From the Sun and Moon to Wormholes and Warp Drive, Key Theories, Discoveries, and Facts about the Universe (The Adams 101 Series)
03Astronomy 101: From the Sun and Moon to Wormholes and Warp Drive, Key Theories, Discoveries, and Facts about the Universe (The Adams 101 Series) -
 04True Anomaly hires former York Space executive as chief operating officer
04True Anomaly hires former York Space executive as chief operating officer -
 05Φsat-2 begins science phase for AI Earth images
05Φsat-2 begins science phase for AI Earth images -
 06Hurricane forecasters are losing 3 key satellites ahead of peak storm season − a meteorologist explains why it matters
06Hurricane forecasters are losing 3 key satellites ahead of peak storm season − a meteorologist explains why it matters -
 07Binary star systems are complex astronomical objects − a new AI approach could pin down their properties quickly
07Binary star systems are complex astronomical objects − a new AI approach could pin down their properties quickly