Now Reading: Genome Sequence And Methylome Of The Extremely Halophilic Bacterium Salinibacter Ruber Strain M31T Isolated From A Crystallizer Pond In Mallorca, Spain
-
01
Genome Sequence And Methylome Of The Extremely Halophilic Bacterium Salinibacter Ruber Strain M31T Isolated From A Crystallizer Pond In Mallorca, Spain
Genome Sequence And Methylome Of The Extremely Halophilic Bacterium Salinibacter Ruber Strain M31T Isolated From A Crystallizer Pond In Mallorca, Spain
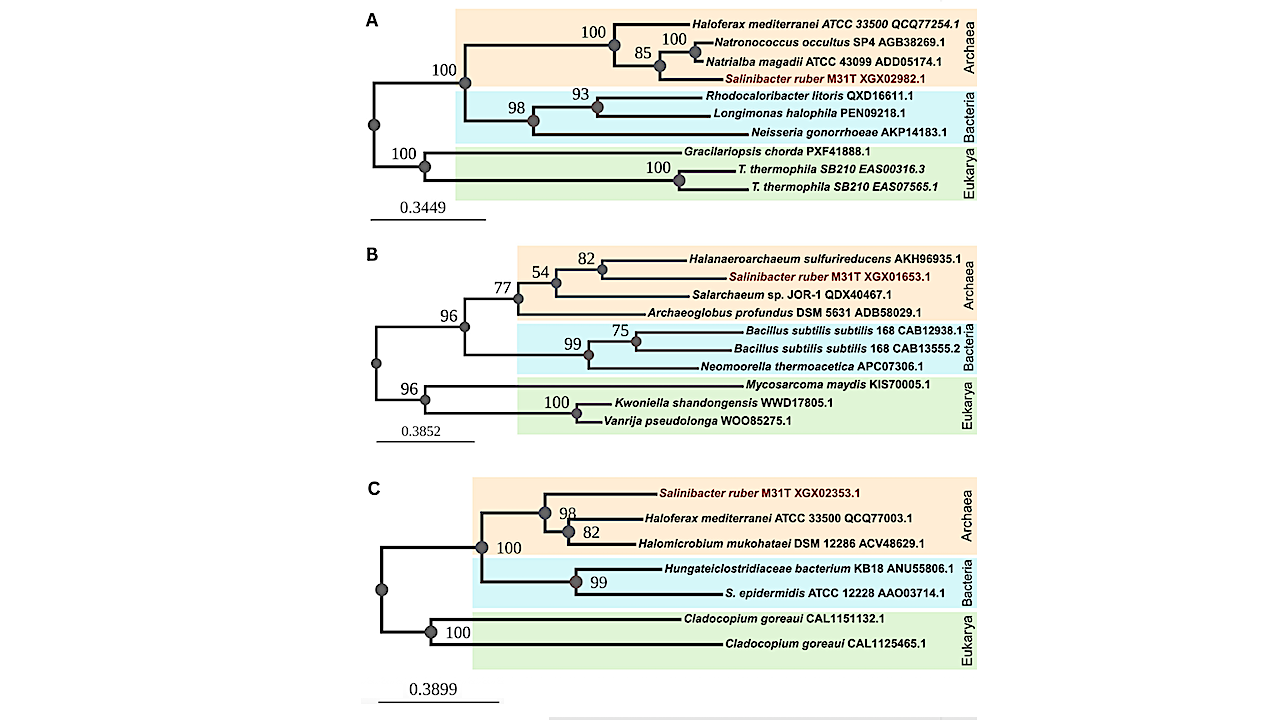
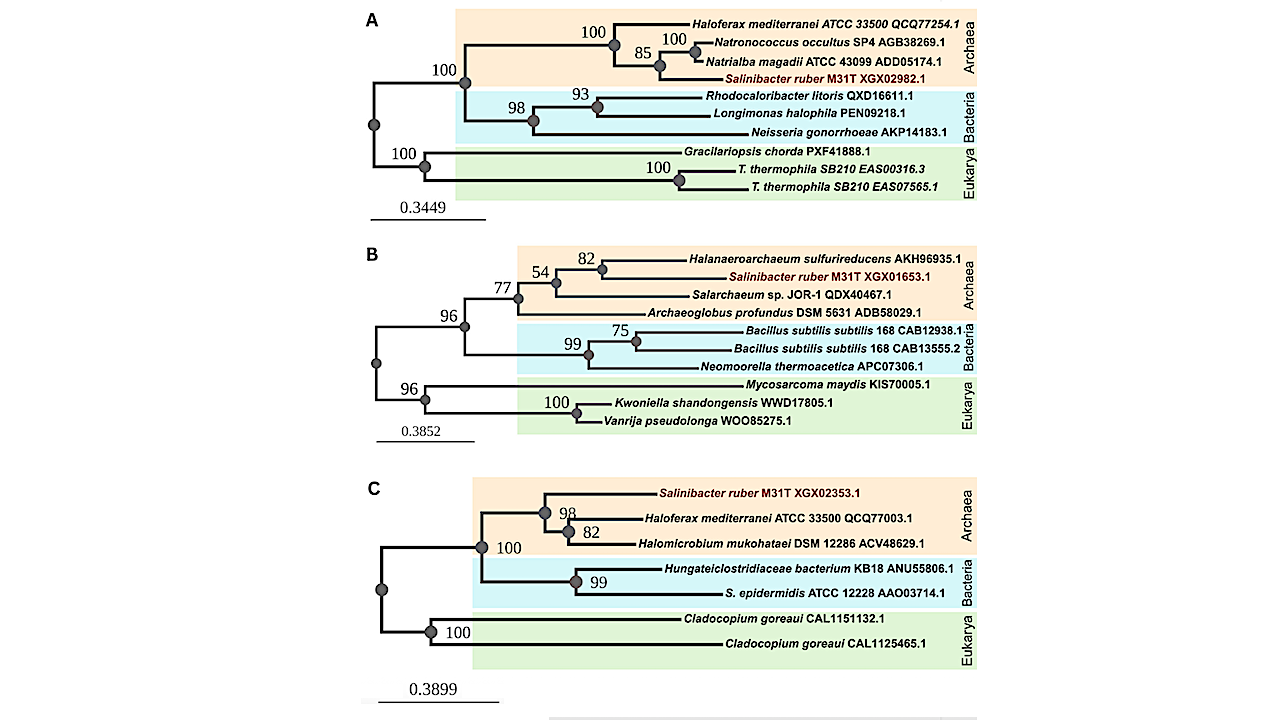
Maximum likelihood phylogenetic trees of the S. ruber M31T-predicted proteins with an archaeal evolutionary history potentially acquired by horizontal gene transfer: (A) sodium-dependent transporter (XGX02982.1); (B) MFS transporter (XGX01653.1); and (C) ABC transporter ATP-binding protein (XGX02353.1). The organism names are followed by the PID of the sequence from NCBI. The S. ruber M31T sequences group with the archaeal sequences (orange) instead of bacterial (blue) or eukaryotic (green) ones. — Environmental Microbiology
Salinibacter ruber strain M31T, an extremely halophilic bacterium, was isolated from a saltern crystallizer pond in Spain. Single-molecule real-time sequencing revealed a 3.6-Mbp genome with a single 3.55-Mbp circular chromosome and a 35.5-kbp plasmid. The highly acidic proteome includes a total of 2,962 proteins, some of which are archaeal-like.
Besides an elevated intracellular concentration of KCl for osmotic balance, it also possesses rhodopsins for phototrophic energy production and displays a high degree of genome plasticity (4).
The gram-negative Salinibacter ruber strain M31T was isolated from a saltern crystallizer pond in Mallorca, Balearic Islands, Spain (GPS: 39.3499° N, 3.0110° E) on September 1999 and was a gift from the American Type Culture Collection (1). Salinibacter ruber M31T cultures were grown in ATCC 2402 medium (5). High-molecular weight genomic DNA was prepared by lysis and extraction with phenol:chloroform, followed by ethanol precipitation as described earlier (6).
Astrobiology,
Stay Informed With the Latest & Most Important News
-
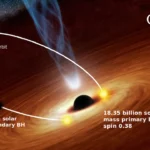 01Two Black Holes Observed Circling Each Other for the First Time
01Two Black Holes Observed Circling Each Other for the First Time -
 02From Polymerization-Enabled Folding and Assembly to Chemical Evolution: Key Processes for Emergence of Functional Polymers in the Origin of Life
02From Polymerization-Enabled Folding and Assembly to Chemical Evolution: Key Processes for Emergence of Functional Polymers in the Origin of Life -
 03Astronomy 101: From the Sun and Moon to Wormholes and Warp Drive, Key Theories, Discoveries, and Facts about the Universe (The Adams 101 Series)
03Astronomy 101: From the Sun and Moon to Wormholes and Warp Drive, Key Theories, Discoveries, and Facts about the Universe (The Adams 101 Series) -
 04True Anomaly hires former York Space executive as chief operating officer
04True Anomaly hires former York Space executive as chief operating officer -
 05Φsat-2 begins science phase for AI Earth images
05Φsat-2 begins science phase for AI Earth images -
 06Hurricane forecasters are losing 3 key satellites ahead of peak storm season − a meteorologist explains why it matters
06Hurricane forecasters are losing 3 key satellites ahead of peak storm season − a meteorologist explains why it matters -
 07Binary star systems are complex astronomical objects − a new AI approach could pin down their properties quickly
07Binary star systems are complex astronomical objects − a new AI approach could pin down their properties quickly