Now Reading: Molecular Complexity Constrained Early Amino Acid Recruitment into the Genetic Code
-
01
Molecular Complexity Constrained Early Amino Acid Recruitment into the Genetic Code
Molecular Complexity Constrained Early Amino Acid Recruitment into the Genetic Code
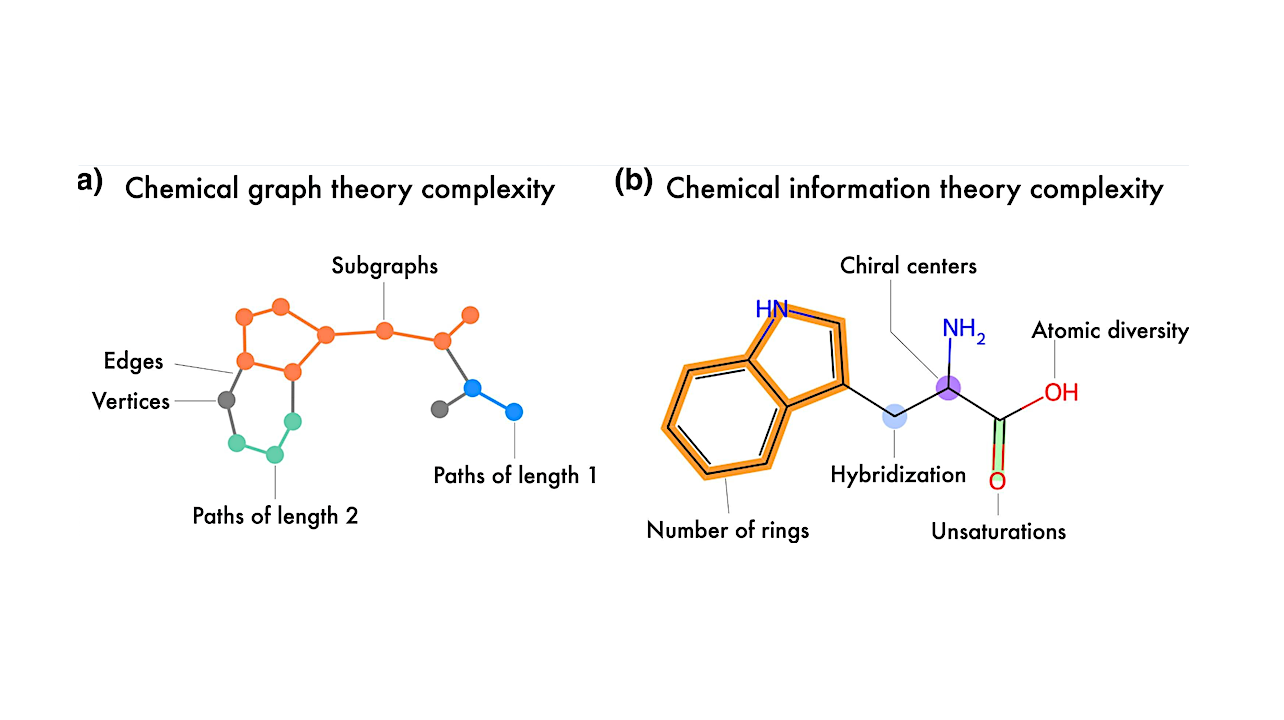
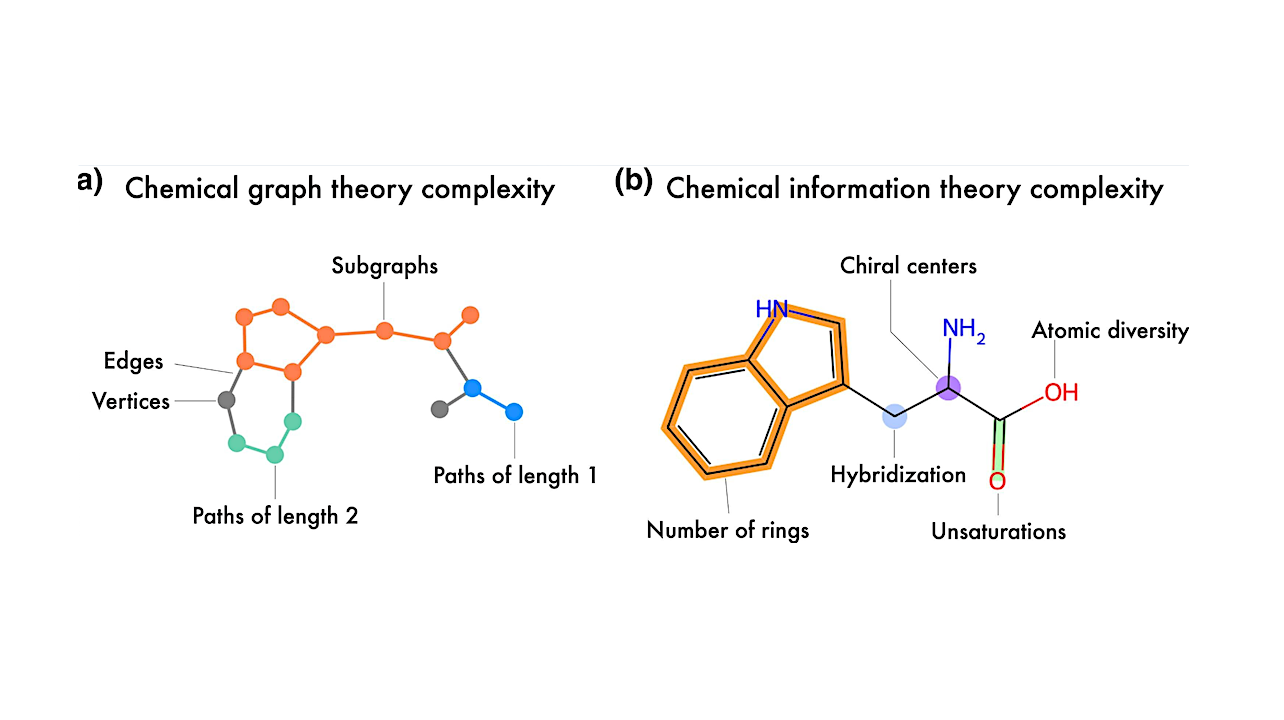
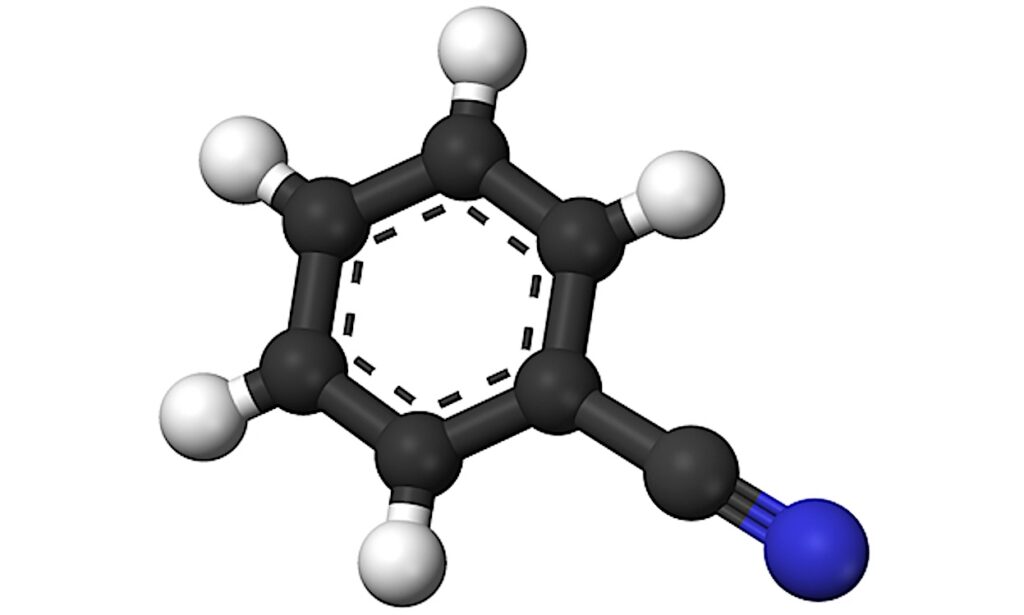
Representation of the two main frameworks used to quantify molecular complexity: (a) graph-theoretic and b) information-theoretic approaches. Tryptophan is depicted as both a graph representation and a molecular structure, illustrating some of the key features captured by each framework. — Genome Biology and Evolution via PubM
Previously proposed chronologies of amino acid incorporation into the genetic code rely on consensus rankings derived from prebiotic synthesis experiments, biosynthetic pathways, or genomic trends.
However, the role of intrinsic molecular properties in shaping amino acid recruitment remains largely underexplored. In this study, we reconstruct a complexity-based amino acid chronology by integrating 16 molecular complexity metrics from chemical graph and information theory.
Unlike approaches influenced by environmental variability, detection biases, or the evolutionary constraints of genome-based chronologies, our method provides a perspective on amino acid incorporation independent of these factors. Instead of imposing a linear ranking, we derive a minimum spanning tree capturing complexity-based relationships between amino acids.
The resulting hierarchy places structurally simple amino acids in basal positions, while biosynthetically complex residues appear later, aligning with existing prebiotic and genomic chronologies. Furthermore, amino acids positioned closer in the complexity space exhibit significantly greater mutational connectivity than expected by chance, suggesting that molecular complexity reflects underlying structural considerations that constrained the genetic code’s evolutionary pathways.
This supports the idea that the code evolved not only to maintain biochemical stability but also to facilitate complexity-preserving substitutions, ensuring smooth adaptive transitions while minimizing energetic cost differences. Additionally, molecular complexity significantly correlates with amino acid enrichment in LUCA’s inferred proteome, reinforcing its role as a fundamental constraint on early protein evolution.
Our approach, rooted in intrinsic molecular properties rather than external contingencies, offers new insights into the constraints shaping the genetic code and expands the scope for identifying universal principles of biochemical evolution.
astrobiology,
Stay Informed With the Latest & Most Important News
Previous Post
Next Post
-
 01Two Black Holes Observed Circling Each Other for the First Time
01Two Black Holes Observed Circling Each Other for the First Time -
 02From Polymerization-Enabled Folding and Assembly to Chemical Evolution: Key Processes for Emergence of Functional Polymers in the Origin of Life
02From Polymerization-Enabled Folding and Assembly to Chemical Evolution: Key Processes for Emergence of Functional Polymers in the Origin of Life -
 03Astronomy 101: From the Sun and Moon to Wormholes and Warp Drive, Key Theories, Discoveries, and Facts about the Universe (The Adams 101 Series)
03Astronomy 101: From the Sun and Moon to Wormholes and Warp Drive, Key Theories, Discoveries, and Facts about the Universe (The Adams 101 Series) -
 04True Anomaly hires former York Space executive as chief operating officer
04True Anomaly hires former York Space executive as chief operating officer -
 05Φsat-2 begins science phase for AI Earth images
05Φsat-2 begins science phase for AI Earth images -
 06Hurricane forecasters are losing 3 key satellites ahead of peak storm season − a meteorologist explains why it matters
06Hurricane forecasters are losing 3 key satellites ahead of peak storm season − a meteorologist explains why it matters -
 07Binary star systems are complex astronomical objects − a new AI approach could pin down their properties quickly
07Binary star systems are complex astronomical objects − a new AI approach could pin down their properties quickly