Now Reading: Sequence Motif Dynamics In RNA Pools
-
01
Sequence Motif Dynamics In RNA Pools
Sequence Motif Dynamics In RNA Pools
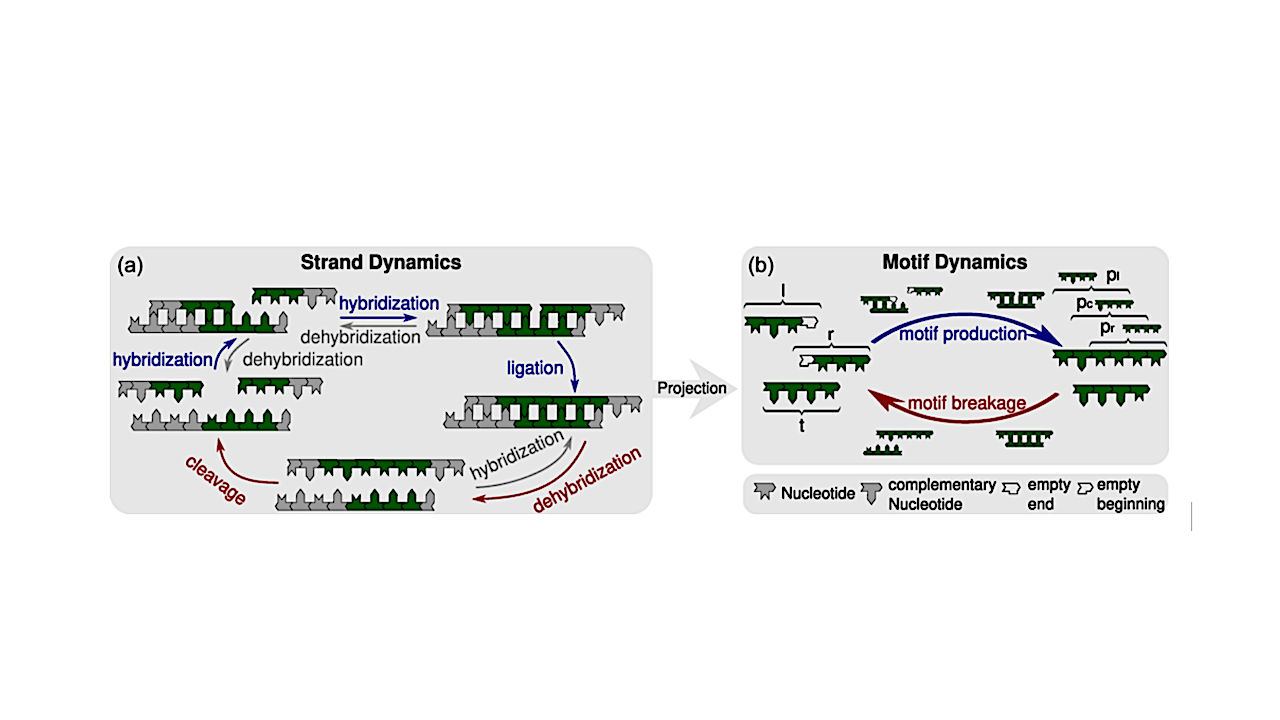
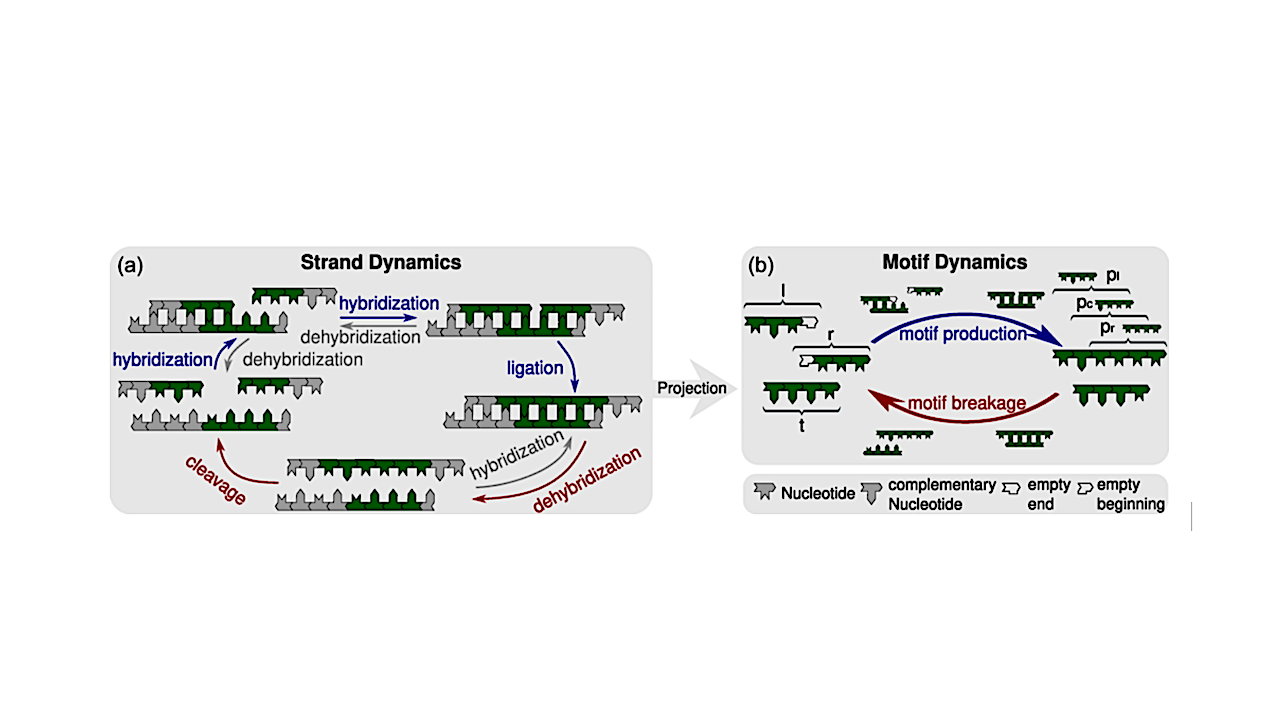
Kinetic processes of the strand-based RNA reactor model [12,13,15] and their projection onto motif kinetics. We consider heteropolymer strands consisting of the two complementary nucleotides X and Y, which could represent RNA or DNA nucleotides. (a) Schematic of the reactions within the strand reactor, including hybridization, dehybridization, templated ligation, and cleavage of oligonucleotide strands. Strands that are not covalently linked are separated by small gaps; four-nucleotide motifs, including 3′ or 5′ ends, are highlighted in green. In templated ligation, a 3′- and a 5′-end motif build a six-nucleotide motif that contains three overlapping four-nucleotide motifs. (b) At the level of sequence motifs, the kinetics are based on motif production and motif breakage reactions. In four-nucleotide motifs, strand ends are marked with 3′ or 5′, respectively. Templated ligation generates longer strands containing additional motifs, thereby contributing to motif production, while strand cleavage leads to to motif breakage. Hybridization is not explicitly represented in the motif-level description; instead, it is implicitly incorporated into the rate constants for motif production and breakage by assuming a hybridization-dehybridization equilibrium and deducing the corresponding dissociation constants. — Physical Review E
In RNA world scenarios, pools of RNA oligomers form strongly interacting, dynamic systems, which enable molecular evolution.
In such pools, RNA oligomers hybridize and dehybridize, ligate, and break, ultimately generating longer RNA molecules, which may fold into catalytically active ribozymes. A key process for the elongation of RNA oligomers is templated ligation, which can occur when two RNA strands are adjacently hybridized onto a template strand.
Detailed simulations of the dynamics in RNA pools involve a large variety of possible sequences and reactions. Here we develop a reduced description of these complex dynamics within the space of sequence motifs. We then explore to what extent our reduced description can capture the behavior of detailed simulations that account for the full dynamics in the space of RNA strands.
Towards this end, we project the dynamics into a motif space, which accounts only for the abundance of all possible four-nucleotide motifs. A system of ordinary differential equations describes the dynamics of those motifs. Its control parameters are effective rate constants for reactions in motif space, which we obtain from the rate constants for the processes underlying the full dynamics in the space of RNA strands.
We find that these reduced motif space dynamics indeed capture important aspects of the informational dynamics of RNA pools in sequence space. This approach could also provide a framework to rationalize and interpret features of the sequence dynamics observed in experimental systems.
Sequence Motif Dynamics In RNA Pools, Physical Review
Astrobiology,
Stay Informed With the Latest & Most Important News
Previous Post
Next Post
-
 01Two Black Holes Observed Circling Each Other for the First Time
01Two Black Holes Observed Circling Each Other for the First Time -
 02From Polymerization-Enabled Folding and Assembly to Chemical Evolution: Key Processes for Emergence of Functional Polymers in the Origin of Life
02From Polymerization-Enabled Folding and Assembly to Chemical Evolution: Key Processes for Emergence of Functional Polymers in the Origin of Life -
 03Astronomy 101: From the Sun and Moon to Wormholes and Warp Drive, Key Theories, Discoveries, and Facts about the Universe (The Adams 101 Series)
03Astronomy 101: From the Sun and Moon to Wormholes and Warp Drive, Key Theories, Discoveries, and Facts about the Universe (The Adams 101 Series) -
 04True Anomaly hires former York Space executive as chief operating officer
04True Anomaly hires former York Space executive as chief operating officer -
 05Φsat-2 begins science phase for AI Earth images
05Φsat-2 begins science phase for AI Earth images -
 06Hurricane forecasters are losing 3 key satellites ahead of peak storm season − a meteorologist explains why it matters
06Hurricane forecasters are losing 3 key satellites ahead of peak storm season − a meteorologist explains why it matters -
 07Binary star systems are complex astronomical objects − a new AI approach could pin down their properties quickly
07Binary star systems are complex astronomical objects − a new AI approach could pin down their properties quickly