Now Reading: Tricorder Tech: Protein Sequencing Advance Offers New Insights Into Life’s Foundations
-
01
Tricorder Tech: Protein Sequencing Advance Offers New Insights Into Life’s Foundations
Tricorder Tech: Protein Sequencing Advance Offers New Insights Into Life’s Foundations
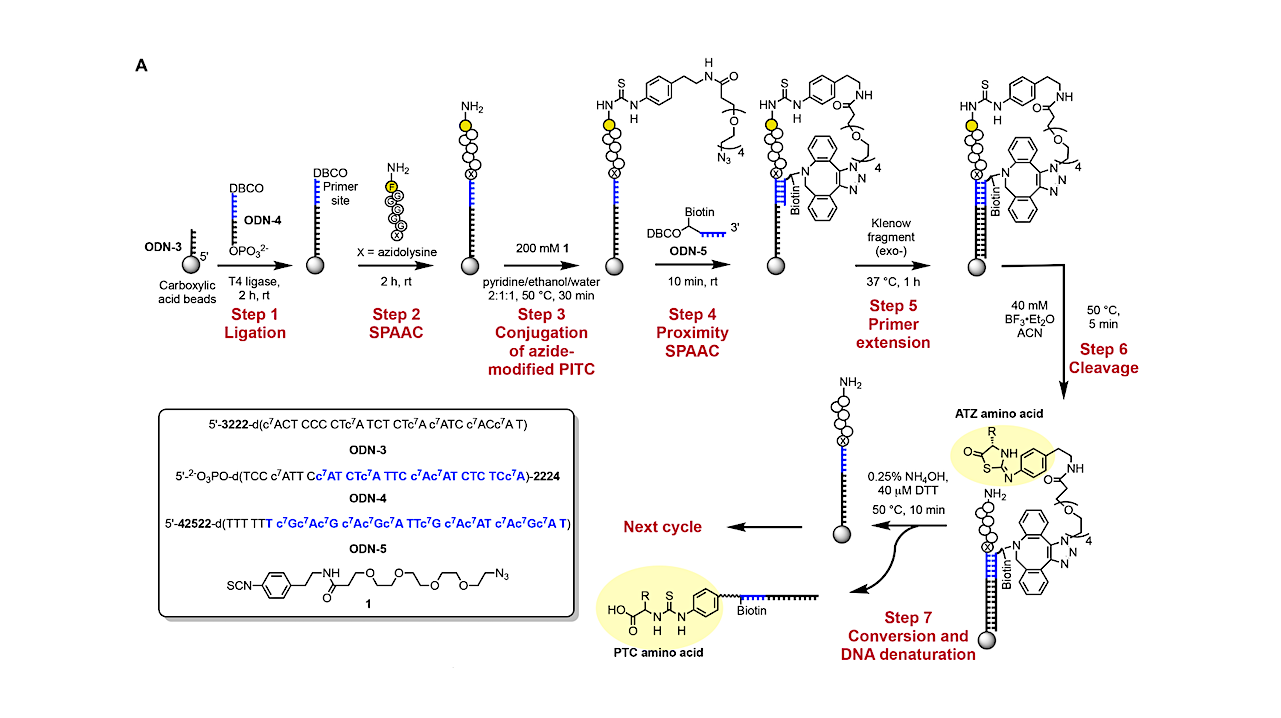

DNA-encoded Edman degradation on magnetic beads. (A) Reaction scheme for barcoding PTC amino acids via DNA-encoded Edman degradation on magnetic beads — Nature Biotechnology
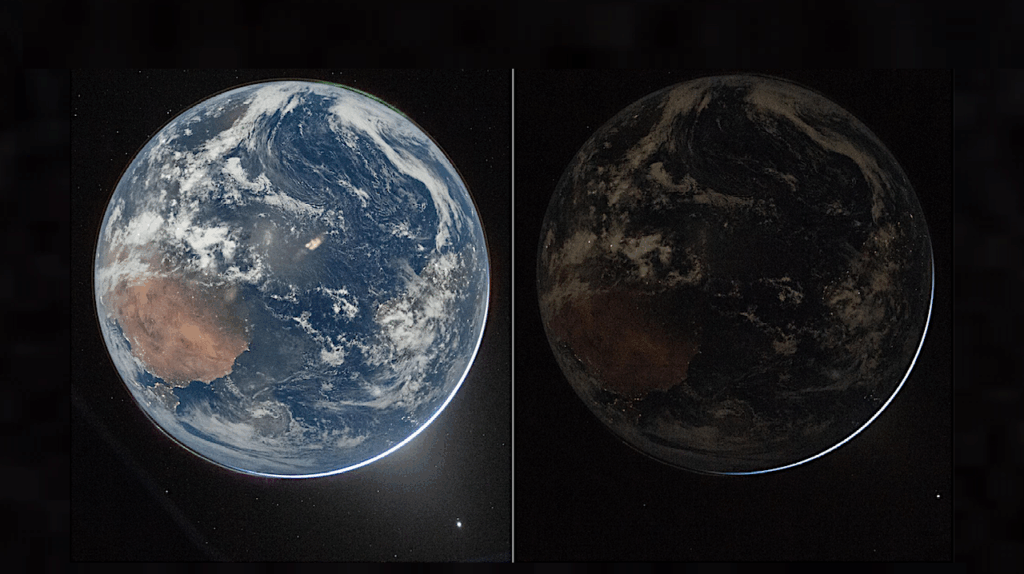
Editor’s note: with recent speculation about the potential habitability of other worlds and understanding the processes whereby other biospheres arise and evolve, thought needs to be given to developing sensor technology to do in situ analysis of samples collected by robotic systems and human crews. With size and power and energy constraints in mind, having the most compact way to quickly analyze a sample or monitor alien life forms will be of great utility to future astrobiology expeditions. Systems such as the one described below are good step in this direction.
Proteins, one of the smallest building blocks of life on Earth, hold promise for answering some of biology’s biggest questions. Consisting of amino acids strung together into peptide chains, these molecules perform much of the work inside living cells. While they execute life’s most essential functions with apparent ease, decoding their precise sequence and structure has long been one of biology’s hardest challenges.
Now, a team led by bioengineers at Stanford University has developed a novel approach to visualize proteins at unprecedented scale and sensitivity. The work, published March 18 in Nature Biotechnology, describes a chemical process that allows researchers to use existing, fast, and low-cost DNA sequencing platforms to decode proteins.
“In nature, proteins are made from DNA. Over the past two decades, our society has created amazing technologies to sequence a lot of DNA really quickly and inexpensively,” said H. Tom Soh, the W. M. Keck Foundation Professor of Electrical Engineering and professor of bioengineering and of radiology in the schools of Engineering and Medicine and senior author of the study. “But unfortunately, we haven’t really made similar progress for sequencing proteins. In this work, we created a technology that can convert protein sequences back into DNA sequences. It’s kind of like running the natural process – in reverse – so that we can leverage powerful DNA sequencing technology that is already available.”

Overview of peptide sequencing via “reverse translation”. In stage 1, peptides are conjugated with peptide barcode DNA sequences immobilized on magnetic beads. A modified Edman degradation reaction is carried out using a PITC molecule modified with an azide group to enable click coupling to a DBCO-modified, biotinylated primer that hybridizes directly to the peptide barcode sequence. This primer is extended to record the peptide barcode before Nterminal amino acid cleavage. In stage 2, the DNA-barcoded amino acids released after cleavage are pulled down by streptavidin (SA) beads via their biotin moiety. These immobilized amino acids are recognized by DNA-barcoded amino acid-specific antibodies, and the DNA-barcoded amino acids are subsequently converted into DNA reporters via proximity extension assay (PEA). In stage 3, the DNA reporters from each cycle are barcoded with cycle number barcodes during adaptor PCR, yielding a DNA sequence that encodes the identity, position, and originating peptide of each amino acid. The final DNA library produced by this process can then be sequenced in a single run, enabling the reconstruction of peptide sequences from the sample. — Nature Biotechnology
The mysteries of proteins
Proteins contain biological information that DNA cannot fully reveal. DNA acts as an instruction manual, carrying the code needed to build cellular machinery. Proteins are that machinery, folding into intricate shapes that determine how cells grow, communicate, respond to immune threats, and malfunction in disease. Despite their central role, scientists have lacked a protein sequencing tool that can match the speed of modern DNA sequencing.
Sequencing proteins has remained notoriously difficult because they are built from 20 different amino acids – far more than the four bases that make up DNA. Those amino acids are also nearly three times smaller than nucleic acids, making them far harder for technologies to reliably detect and distinguish.
“What really is different is how much from the same sample we can see,” said Liwei Zheng, research engineer at Stanford and first author of the study. “With mass spectrometry, you’re shooting 1 billion to 10 billion protein molecules and see, typically, a million molecules out of it. With our method, you can potentially see 1,000 times that amount.” Zheng hopes to see more rare types of proteins brought to light using this new sequencing.
Binding (and finding) the solution
The team’s new chemistry tackles this problem by tagging individual amino acids within a peptide using DNA barcodes specific to each peptide molecule. Antibodies and cycle barcodes, known as synthetic DNA, then help encode the peptide’s identity and mark its position, respectively. Once encoded, routine DNA sequencing can reveal the identity and location of the amino acid within the peptide chain.
“This is the breakthrough,” said Soh. “We can sequence individual proteins at a single molecule level, requiring very little sample that can get us to individual cells.”
This sensitivity could enable studies of cellular diversity at nanoscopic resolution, shedding light on why seemingly identical cells behave differently in response to diseases like cancer. It also opens up the possibility of studying rare proteins that are difficult to detect in bulk samples, including molecules that may drive disease progression but exist only in tiny quantities.
Soh points to immunotherapy as one example: treatments like CAR-T cell therapy have revolutionized cancer care by modifying patients’ own immune cells to recognize and fight back against cancer cells. A big unknown is why it only works for certain types of cancers and certain types of cells.
By isolating immune cells that positively respond to treatments with other cells that don’t, using a single-protein level analysis, scientists can more clearly understand why certain treatments work and how they can be enhanced to maximize efficiency across individual patients.
From bench to button
The breakthrough has already sparked commercial interest and is licensed with the intent of turning the laboratory process into a user-friendly instrument. The goal is to eventually enable researchers to “put in a sample, press a button, and have it go,” said Soh, much like pressing a button to run a DNA sequencer today.
While the method is still in its early stages and will require optimization before widespread adoption, its ability to potentially scale billions of protein molecules from thousands of cells sets it apart from previous attempts at direct protein sequencing.
“Once you convert everything to DNA, you can think about all the naturally evolved machinery that can manipulate DNA – lengthening DNA, copying DNA – all these things will become possible for processing protein sequences,” said Zheng.
If successful, the team’s “reverse translation” approach could become a new cornerstone of molecular biology, giving scientists a long-missing tool to read the molecules that do life’s work, one cell at a time.
Acknowledgements:
Additional co-authors of this paper are postdoctoral fellow Yujia Sun, graduate student Linus Hein, and researcher Michael Eisenstein. Soh is also a member of Stanford Bio-X, the Wu Tsai Human Performance Alliance, the Maternal & Child Health Research Institute (MCHRI), the Stanford Cancer Institute, and the Wu Tsai Neurosciences Institute.
Zheng, Sun, and Soh are listed as coinventors on a pending patent application related to this work filed at the U.S. Patent and Trademark Office.
This work was funded by the Helmsley Charitable Trust and the Wellcome Leap SAVE program. It used the Vincent Coates Foundation Mass Spectrometry Laboratory and Stanford University Mass Spectrometry’s Bruker Microflex MALDI TOF mass spectrometer and Thermo Exploris 240 LC/MS system, purchased with funding from Stanford C-ShaRP.
Astrobiology,
Stay Informed With the Latest & Most Important News
Previous Post
Next Post
-
 01Two Black Holes Observed Circling Each Other for the First Time
01Two Black Holes Observed Circling Each Other for the First Time -
 02From Polymerization-Enabled Folding and Assembly to Chemical Evolution: Key Processes for Emergence of Functional Polymers in the Origin of Life
02From Polymerization-Enabled Folding and Assembly to Chemical Evolution: Key Processes for Emergence of Functional Polymers in the Origin of Life -
 03Astronomy 101: From the Sun and Moon to Wormholes and Warp Drive, Key Theories, Discoveries, and Facts about the Universe (The Adams 101 Series)
03Astronomy 101: From the Sun and Moon to Wormholes and Warp Drive, Key Theories, Discoveries, and Facts about the Universe (The Adams 101 Series) -
 04True Anomaly hires former York Space executive as chief operating officer
04True Anomaly hires former York Space executive as chief operating officer -
 05Φsat-2 begins science phase for AI Earth images
05Φsat-2 begins science phase for AI Earth images -
 06Hurricane forecasters are losing 3 key satellites ahead of peak storm season − a meteorologist explains why it matters
06Hurricane forecasters are losing 3 key satellites ahead of peak storm season − a meteorologist explains why it matters -
 07Binary star systems are complex astronomical objects − a new AI approach could pin down their properties quickly
07Binary star systems are complex astronomical objects − a new AI approach could pin down their properties quickly