Now Reading: Fungal Species Discovered With The Ability To Freeze Water
-
01
Fungal Species Discovered With The Ability To Freeze Water
Fungal Species Discovered With The Ability To Freeze Water
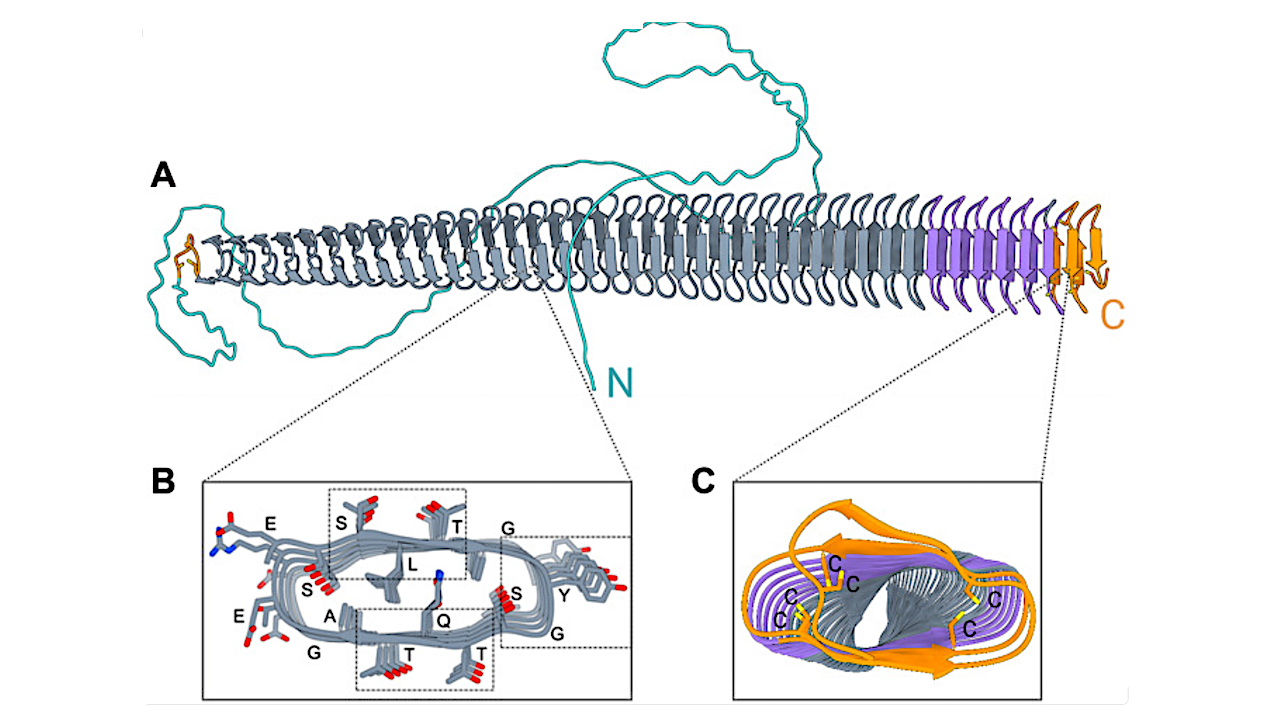
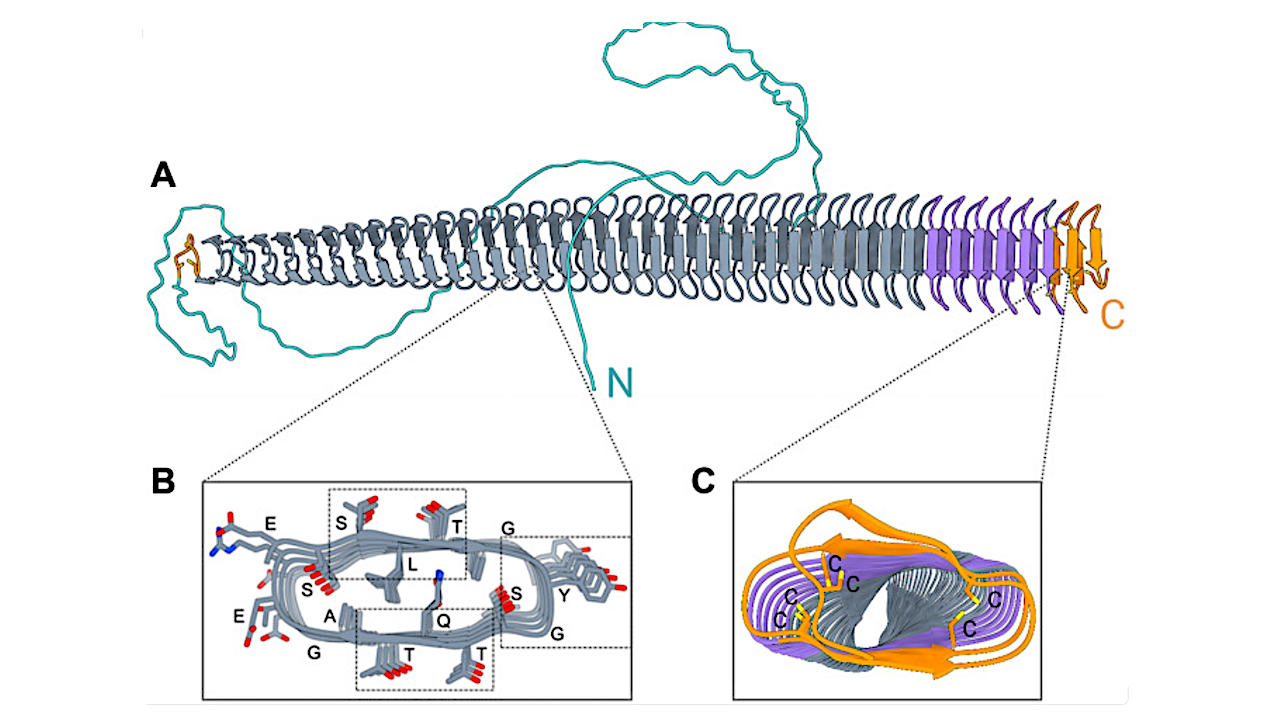
(A) The model of EnINpro is colored by different domains with arrows representing β strands. The model highlights a β-solenoid fold (gray, purple) adjacent to the disulfide capping motif (orange). (B) Cross section through the central domain region. Residues are identified by their one-letter codes. Boxes indicated the location of the characteristic TQT, SLT, and YGS motifs. (C) Cross section of the central domain’s capping structure, where adjacent cysteine residues can form disulfide bonds that likely stabilize the β-solenoid fold. — Science Advances via PubMed
Ice-nucleating proteins (INpros) catalyze ice formation at high subzero temperatures, with major biological and environmental implications.
While bacterial INpros have been structurally characterized, their counterparts in other organisms have remained largely unknown. Here, we identify membrane-independent proteins in fungi of the Mortierellaceae family that promote ice formation with high efficiency. These proteins are predicted to adopt β-solenoid folds and multimerize to form extended ice-binding surfaces, exhibiting mechanistic parallels with bacterial INpros.
Structural modeling, phylogenetic analysis, and heterologous gene expression leading to ice nucleation in Escherichia coli and Saccharomyces cerevisiae show that the fungal INpros are encoded by orthologs of the bacterial InaZ gene, which was likely acquired by a fungal ancestor through horizontal gene transfer.
The discovery of cell-free fungal INpros provides tools for innovative freezing applications and reveals biophysical constraints on ice nucleation across life.
A previously unrecognized class of fungal ice-nucleating proteins with bacterial ancestry, Science Advances via PubMed (open access)
Astrobiology,
Stay Informed With the Latest & Most Important News
-
 01Two Black Holes Observed Circling Each Other for the First Time
01Two Black Holes Observed Circling Each Other for the First Time -
 02From Polymerization-Enabled Folding and Assembly to Chemical Evolution: Key Processes for Emergence of Functional Polymers in the Origin of Life
02From Polymerization-Enabled Folding and Assembly to Chemical Evolution: Key Processes for Emergence of Functional Polymers in the Origin of Life -
 03Astronomy 101: From the Sun and Moon to Wormholes and Warp Drive, Key Theories, Discoveries, and Facts about the Universe (The Adams 101 Series)
03Astronomy 101: From the Sun and Moon to Wormholes and Warp Drive, Key Theories, Discoveries, and Facts about the Universe (The Adams 101 Series) -
 04True Anomaly hires former York Space executive as chief operating officer
04True Anomaly hires former York Space executive as chief operating officer -
 05Φsat-2 begins science phase for AI Earth images
05Φsat-2 begins science phase for AI Earth images -
 06Hurricane forecasters are losing 3 key satellites ahead of peak storm season − a meteorologist explains why it matters
06Hurricane forecasters are losing 3 key satellites ahead of peak storm season − a meteorologist explains why it matters -
 07Binary star systems are complex astronomical objects − a new AI approach could pin down their properties quickly
07Binary star systems are complex astronomical objects − a new AI approach could pin down their properties quickly