Now Reading: Computational Chemistry And Supercomputers To Help Understand The Mechanisms Of Life
-
01
Computational Chemistry And Supercomputers To Help Understand The Mechanisms Of Life
Computational Chemistry And Supercomputers To Help Understand The Mechanisms Of Life
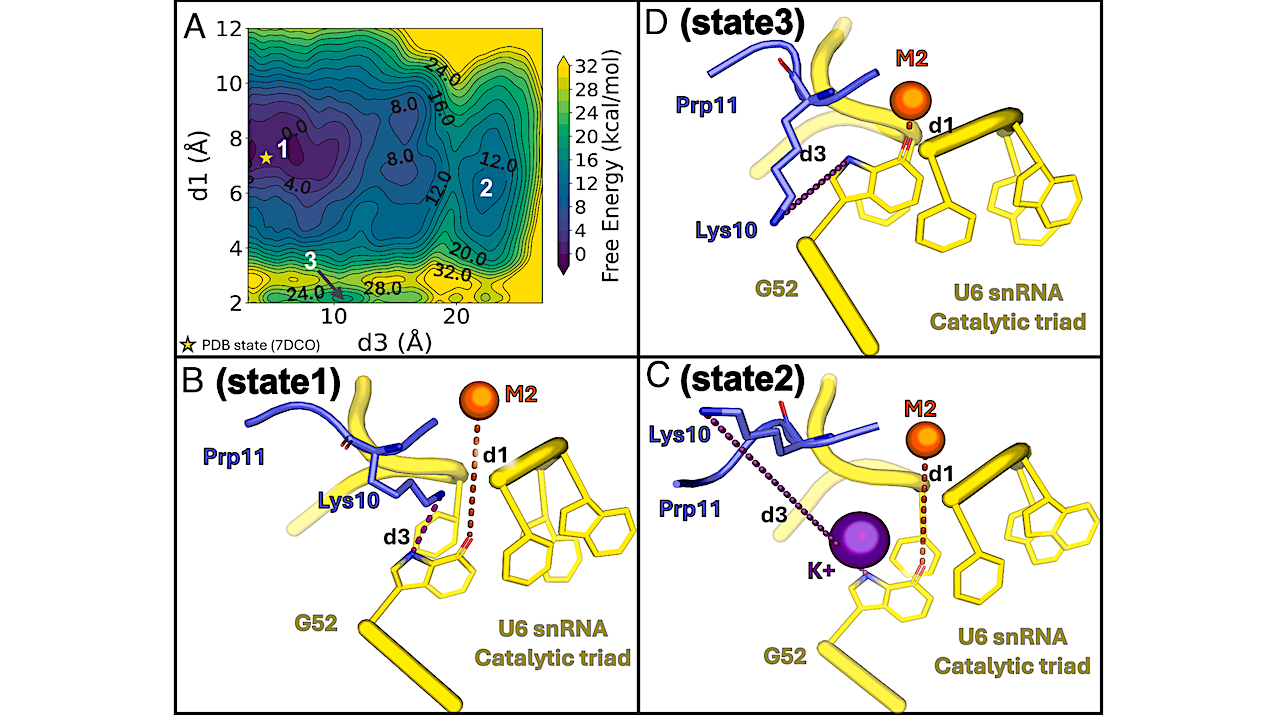
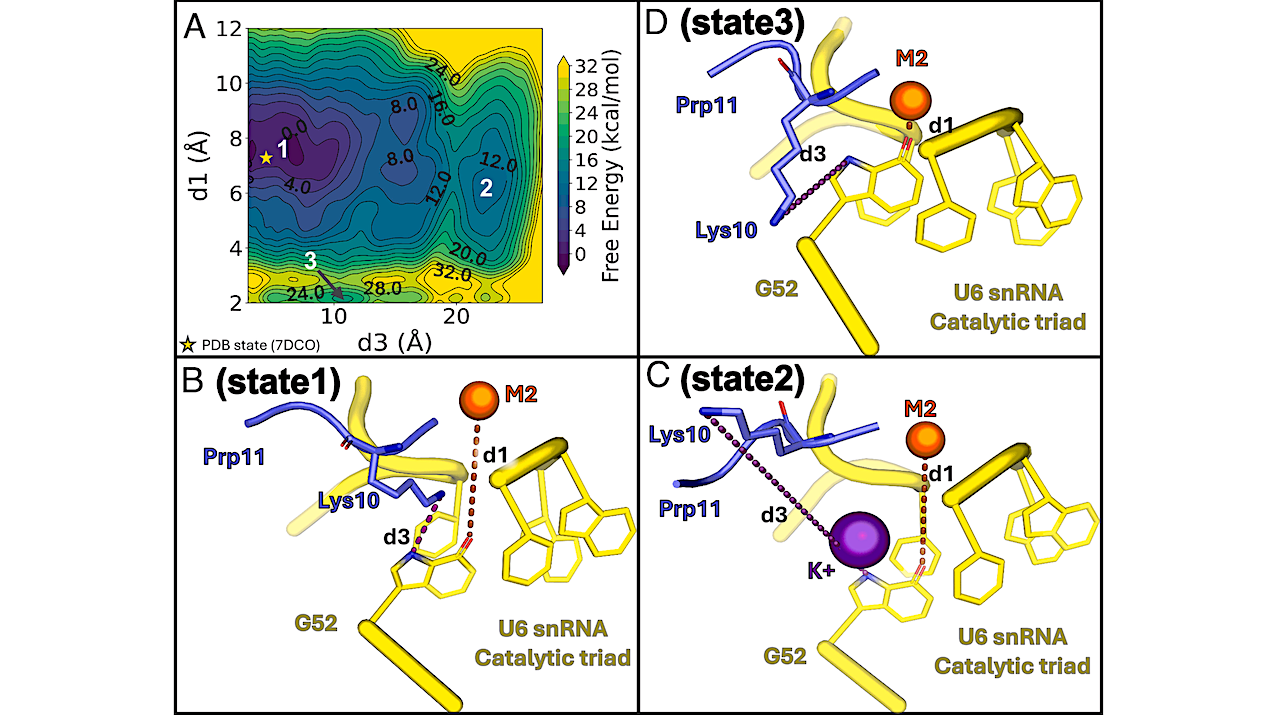
Energetics associated with the occupancy of the K1 site and the position of the M2 ion. (A) Free energy surface of the concerted motions of Prp11 Lys10 and M2, with the initial PDB conformation marked by a yellow star. (B–D) Representative structures of three metastable states from metadynamics simulations. State 1 (B) represents a PDB-like arrangement where Lys10 (slate blue sticks) remains at the K1 site, and M2 (yellow sphere) is properly positioned at the U6 snRNA triple-helix (yellow cartoon). State 2 (C): Lys10 leaves the K1 site, allowing a K+ ion to occupy it and stabilize M2 in place. State 3 (D): M2 directly coordinates G52, losing its catalytic position and partially occupying the K1 site. — PNAS
A new study from the Italian Institute of Technology (IIT), in collaboration with Uppsala University (Sweden) and AstraZeneca, shows how computational chemistry and supercomputers can help scientists better understand the fundamental mechanisms of life, specifically those of human cells.
This research was conducted by the Molecular Modeling and Drug Discovery Unit, led by Marco De Vivo at IIT in Genoa, and was published in the scientific journal Proceedings of the National Academy of Sciences (PNAS). The study focused on splicing, an essential process that enables cells to read genes and function properly, by simulating the functional dynamics of a very large and realistic model system (~2 million atoms).
The results provide a more precise understanding of splicing and help explain previously difficult-to-interpret data on splicing. This research could support the development of molecules capable of controlling this process, paving the way for new treatments for diseases such as cancer and neurodegenerative disorders.
Cells work properly thanks to gene expression, the process through which the information in DNA is copied into RNA and then used by the cell. We can think of DNA as a large instruction manual, and gene expression as the system that allows the cell to choose and use the instructions it needs at a given time.
An important step in this process is splicing. During splicing, RNA molecules are reorganized so they can work correctly. This process is controlled by a large, complex structure called the spliceosome, which is composed of many proteins and RNA molecules. Because the spliceosome is very dynamic and complex, it has always been difficult to study in detail.
To better understand it, IIT researchers, with PhD student Gianfranco Martino as the first author of the study, used advanced computer simulations, mainly run on IIT’s supercomputer, “Franklin” (named after Rosalind Franklin, the British scientist who contributed to the discovery of the structure of DNA). These simulations enable scientists to recreate how molecules move and interact over time, in accordance with the laws of physics. This allows observation of biological processes in much greater detail than with experimental methods alone.
Thanks to Franklin’s computing power (powered by more than 360 GPU processors), the researchers were able to study the spliceosome at the atomic level and observe how it changes shape during its activity. This was a major technical challenge: the simulation included about 2 million atoms, far more than traditional simulations, which usually involve between 200,000 and 500,000 atoms.
Until now, scientists only had static snapshots of the spliceosome, images showing what it looks like at different moments. What was missing was an understanding of how it moves from one state to another in a dynamic way.
The new simulations developed at IIT made it possible to observe this process. The researchers discovered that the movements follow a precise and controlled sequence, which is essential for the spliceosome to function correctly. The model also helped explain some experimental data that had previously been difficult to interpret.
“These results show that computational chemistry and molecular simulations, made possible by supercomputers, can strongly support experimental biology when studying very complex systems. Understanding exactly how the spliceosome works is an essential step toward developing new targeted drugs” says Marco De Vivo, Principal Investigator of the Molecular Modeling and Drug Discovery unit at IIT.
“The collaboration between my group, which studies RNA experimentally, and Dr. De Vivo’s team, which specializes in computational drug discovery, is key to moving faster toward our ultimate goal: discovering new therapies” says Marco Marcia, Associate Professor at Uppsala University in Sweden and AIRC-funded visiting PI at IIT.
The next step for the research team will be to improve molecules that have already been identified as capable of regulating the spliceosome’s activity. This could lead to new treatments for diseases in which this process does not work correctly, offering promising opportunities for future medicine.
The research was supported by the Italian AIRC Foundation for Cancer Research and by the European High Performance Computing Joint Undertaking.
Controlled dynamic remodeling of the spliceosome active site enables the first step of splicing, PNAS (open access)
Astrobiology, genomics, evolution,
Stay Informed With the Latest & Most Important News
Previous Post
Next Post
-
 01Two Black Holes Observed Circling Each Other for the First Time
01Two Black Holes Observed Circling Each Other for the First Time -
 02From Polymerization-Enabled Folding and Assembly to Chemical Evolution: Key Processes for Emergence of Functional Polymers in the Origin of Life
02From Polymerization-Enabled Folding and Assembly to Chemical Evolution: Key Processes for Emergence of Functional Polymers in the Origin of Life -
 03Astronomy 101: From the Sun and Moon to Wormholes and Warp Drive, Key Theories, Discoveries, and Facts about the Universe (The Adams 101 Series)
03Astronomy 101: From the Sun and Moon to Wormholes and Warp Drive, Key Theories, Discoveries, and Facts about the Universe (The Adams 101 Series) -
 04True Anomaly hires former York Space executive as chief operating officer
04True Anomaly hires former York Space executive as chief operating officer -
 05Φsat-2 begins science phase for AI Earth images
05Φsat-2 begins science phase for AI Earth images -
 06Hurricane forecasters are losing 3 key satellites ahead of peak storm season − a meteorologist explains why it matters
06Hurricane forecasters are losing 3 key satellites ahead of peak storm season − a meteorologist explains why it matters -
 07Binary star systems are complex astronomical objects − a new AI approach could pin down their properties quickly
07Binary star systems are complex astronomical objects − a new AI approach could pin down their properties quickly