Now Reading: A Ribozyme Mass Extinction At The RNA-cellular Boundary And Its Potential Imprint On The Genetic Code
-
01
A Ribozyme Mass Extinction At The RNA-cellular Boundary And Its Potential Imprint On The Genetic Code
A Ribozyme Mass Extinction At The RNA-cellular Boundary And Its Potential Imprint On The Genetic Code
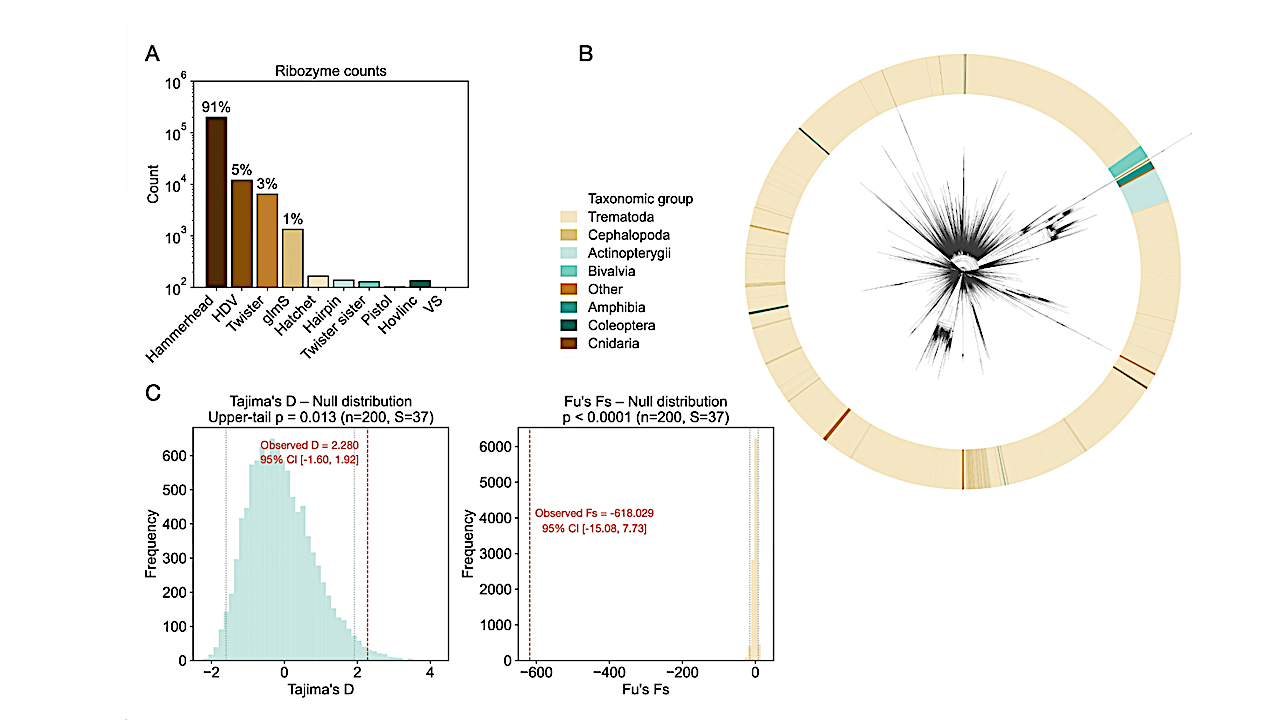

Evidence for ribozyme mass extinction. (A) Abundance distribution across all ten known small self-cleaving ribozyme families, showing the hammerhead’s 91% dominance, a pattern characteristic of disaster taxa following mass extinction events. (B) Circular phylogeny of 80,548 unique hammerhead sequences colored by host taxonomic group. The star-like topology is consistent with post-bottleneck radiation, and the interspersion of distantly related host phyla around the tree reflects repeated independent colonization events rather than co-speciation, consistent with horizontal transfer of a single dominant survivor lineage. (C) Null distribution tests for Tajima’s D (left) and Fu’s Fs (right), each based on 10,000 coalescent replicates (n = 200, S = 40). Blue and orange histograms show the null distributions under neutral expectations; red dashed lines indicate the observed values (D = +2.28, upper-tail p = 0.013; Fs = −618.0, p 0.0001). Observed D falls in the extreme upper tail (98.7% of null replicates below observed), indicating deep hierarchical structure; observed Fs falls far below any null replicate, indicating massive haplotype excess. The discordance between positive D and extremely negative Fs is diagnostic of ancient divergence preserved through a bottleneck followed by rapid within-lineage expansion. Full-sample values (n = 5,036) are D = +3.79, Fs = −690.8. — biorxiv.org
The transition from the RNA world to cellular life remains one of the least understood events in the history of biology.
This study proposes that geochemical changes approximately 3.9-3.8 billion years ago triggered a mass extinction of ribozymes – possibly the first extinction event on Earth – and that the genetic code bears the fingerprints of its survivors.
The argument draws on parallels with animal mass extinctions, where generalist species survive, dominate post-extinction ecosystems as disaster taxa, and shape subsequent evolutionary trajectories. Among small self-cleaving ribozymes, the hammerhead ribozyme accounts for approximately 91% of all ∼221,000 known sequences and is the only ribozyme that is found across all kingdoms of life.
The phylogeny of hammerhead ribozymes exhibits a star-like topology consistent with rapid post-bottleneck expansion, and its small size, ability to tolerate diverse chemical conditions, and broad substrate specificity suggest it was a resilient generalist feeder fitting as a disaster taxon.
Other families of ribozymes possibly survived in small, taxonomically isolated populations resembling relict species in ecological refugia. It is suggested that the body plans of surviving ribozymes seeded the primitive processes that would later become the genetic code, for example with RNA-degrading trinucleotides becoming stop codons, partitioning the trinucleotide space into signals of termination and translation.
This hypothesis proposes a reframing of the origin of the genetic code in part as an ecological legacy rather than a purely chemical inevitability.
A ribozyme mass extinction at the RNA-cellular boundary and its potential imprint on the genetic code, biorxiv.org
Astrobiology, Genomics,
Stay Informed With the Latest & Most Important News
Previous Post
Next Post
-
 01Two Black Holes Observed Circling Each Other for the First Time
01Two Black Holes Observed Circling Each Other for the First Time -
 02From Polymerization-Enabled Folding and Assembly to Chemical Evolution: Key Processes for Emergence of Functional Polymers in the Origin of Life
02From Polymerization-Enabled Folding and Assembly to Chemical Evolution: Key Processes for Emergence of Functional Polymers in the Origin of Life -
 03Astronomy 101: From the Sun and Moon to Wormholes and Warp Drive, Key Theories, Discoveries, and Facts about the Universe (The Adams 101 Series)
03Astronomy 101: From the Sun and Moon to Wormholes and Warp Drive, Key Theories, Discoveries, and Facts about the Universe (The Adams 101 Series) -
 04True Anomaly hires former York Space executive as chief operating officer
04True Anomaly hires former York Space executive as chief operating officer -
 05Φsat-2 begins science phase for AI Earth images
05Φsat-2 begins science phase for AI Earth images -
 06Hurricane forecasters are losing 3 key satellites ahead of peak storm season − a meteorologist explains why it matters
06Hurricane forecasters are losing 3 key satellites ahead of peak storm season − a meteorologist explains why it matters -
 07Binary star systems are complex astronomical objects − a new AI approach could pin down their properties quickly
07Binary star systems are complex astronomical objects − a new AI approach could pin down their properties quickly