Now Reading: New Method Sharpens The Search For Alien Biology
-
01
New Method Sharpens The Search For Alien Biology
New Method Sharpens The Search For Alien Biology
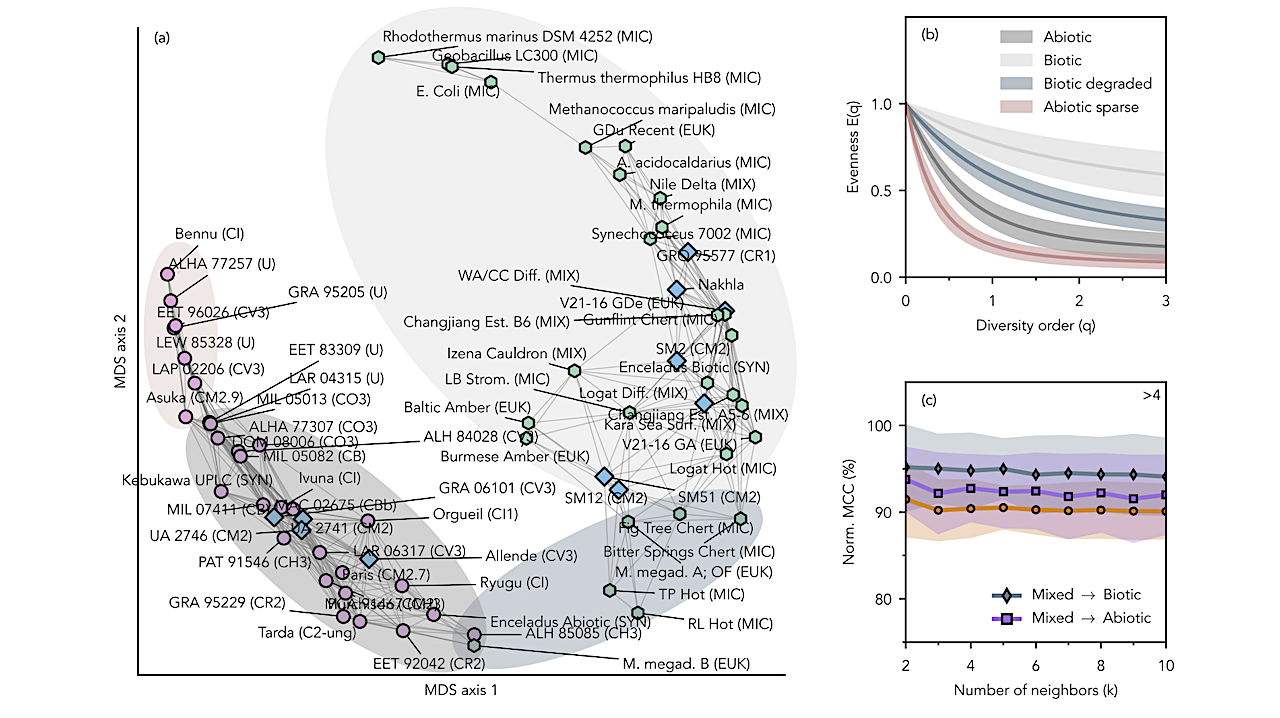
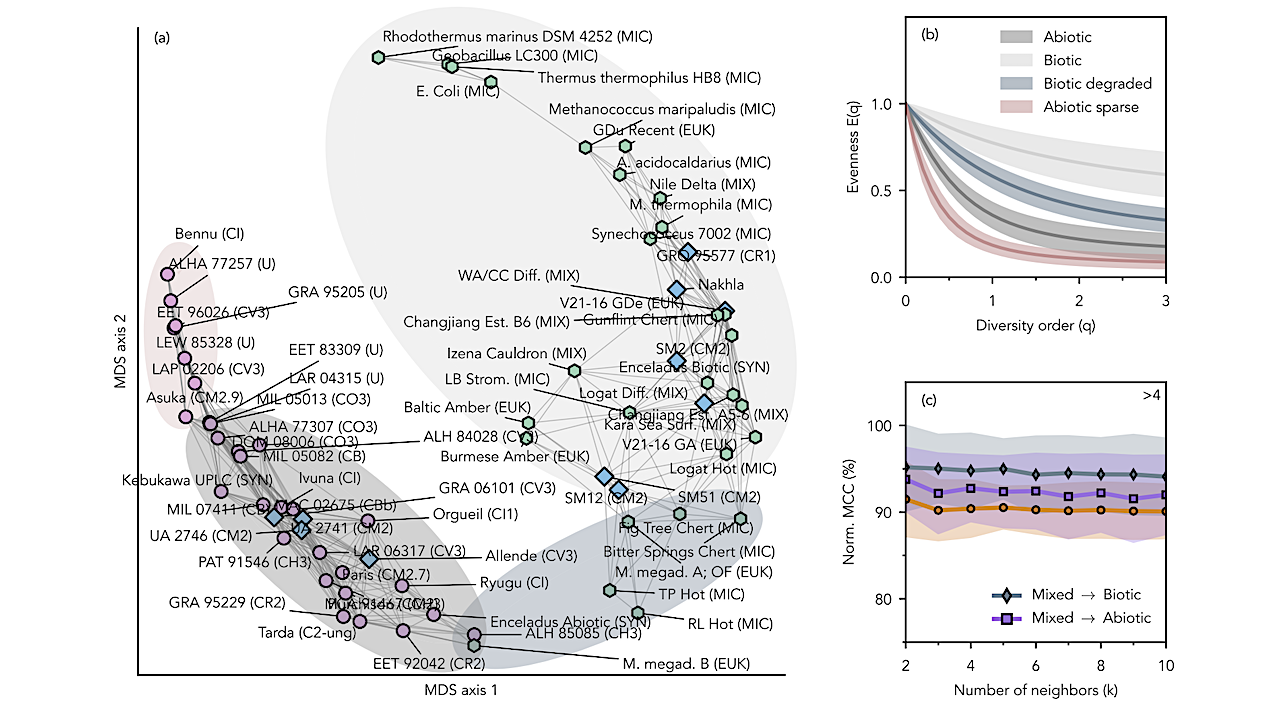
Dissimilarity analysis of evenness curves for amino-acid assemblages. (a) Mul- tidimensional Scaling (MDS) projection of pairwise dissimilarities between evenness curves, E(q). Each point represents a sample; distances increase with statistical separation. Edges connect sam- ples to the 25th percentile of their nearest neighbors. Markers denote inferred origin: biotic (green hexagons), abiotic (pink circles), and mixed (blue diamonds). (b) Evenness-curve distributions for four sample groups. Solid lines indicate group means; shaded regions denote one standard deviation. Each color represents the distribution of samples contained within the shaded regions of the same color in panel (a). (c) Classification performance of sample origin using k-Nearest- Neighbors (kNN) applied to pairwise dissimilarities projected onto the first two MDS axes. Three labeling schemes are evaluated: all three groups retained, and two alternatives in which the mixed group is assigned to either the biotic or abiotic class. Accuracy is reported as the normalized Matthews Correlation Coefficient (MCC), where 50% corresponds to random assignment and 100% to perfect classification. Uncertainty is estimated by bootstrapping class-balanced subsam- ples and recomputing kNN accuracy for each subsample. The value shown at the top right denotes the smallest permutation-based z-score across all k. It measures the deviation of the observed mean MCC from the mean under permuted class labels, using the most conservative value across the three labeling schemes. -— astro-ph.EP
For decades, the search for life beyond Earth has revolved around a key question: What molecules should scientists be looking for on other planets or moons?
A new study, published in Nature Astronomy, suggests the more revealing clue may not be the molecules themselves, but the hidden order connecting them.
“We’re showing that life does not only produce molecules,” said Fabian Klenner, UC Riverside assistant professor of planetary sciences and co-author of the study. “Life also produces an organizational principle that we can see by applying statistics.”
The researchers found that amino acids are consistently more diverse and more evenly distributed in a material sample created by a living thing than those found in abiotic or nonliving things. In contrast, the pattern reverses for fatty acids: abiotically produced fatty acids are distributed more evenly than those produced by biological processes.
This study is the first to demonstrate that this fundamental principle of life can be detected using a statistical approach that does not rely on any one special instrument. Instead, it may be possible to find this pattern in data collected by instruments already aboard current and planned space missions.
The work arrives as planetary exploration enters a new phase in which longstanding questions about the origin of life and its prevalence in the universe may finally become testable with real observational data. Missions to Mars, Europa, Enceladus, and other worlds are returning increasingly sophisticated measurements of organic chemistry. Yet interpreting those measurements remains difficult.
Many compounds central to terrestrial biology, including amino acids and fatty acids, can also form through nonbiological processes. They have been detected in meteorites and synthesized in laboratory experiments designed to mimic conditions in space. Finding such molecules alone is not enough to claim evidence of life.
“Astrobiology is fundamentally a forensic science,” said Gideon Yoffe, postdoctoral researcher at the Weizmann Institute of Science in Israel and first author of the study. “We’re trying to infer processes from incomplete clues, often with very limited data collected by missions that are extraordinarily expensive and infrequent.”
The researchers approached the problem with a statistical framework borrowed from ecology, where scientists quantify biodiversity by measuring two properties: richness, or how many species are present, and evenness, or how uniformly they are distributed. Yoffe first encountered the approach while completing doctoral work in statistics and data science, where diversity metrics were used to uncover patterns in complex datasets, including studies of ancient human cultures.

Dissimilarity analysis of evenness curves of amino acid assemblages. (a): Multidimensional Scaling (MDS) projection of dissimilarities between evenness curves, E(q). Points represent samples; distances between samples grow with dissimilarity. Edges connect samples to the 25th percentile of their nearest neighbors. Markers indicate inferred origin: biotic (green hexagons), abiotic (pink circles), and mixed (blue diamonds). (b): Evenness curve distributions of four distinct sample groups. Solid lines represent mean values, and filled areas represent the one standard deviation interval. (c): Predictive power of a sample’s origin through k-Nearest-Neighbors (kNN) classification, applied to pairwise distances between samples projected onto two MDS axes. Accuracy is quantified using a normalized Matthews Correlation Coefficient (MCC), where 50% corresponds to random assignment and 100% indicates perfect classification. Uncertainty was estimated using multiple initializations of the MDS projection. -— astro-ph.EP
The team applied the same logic to extraterrestrial chemistry.
Using approximately 100 existing datasets, the researchers analyzed amino acids and fatty acids from microbes, soils, fossils, meteorites, asteroids, and synthetic laboratory samples. Biological samples repeatedly exhibited distinct organizational patterns that separated them from nonliving chemistry.
What surprised the researchers most was the method’s strength despite its simplicity.
Looking at the samples in this way, the researchers were consistently able to separate biological and abiotic samples with striking reliability. In addition, they were also able to see that biologically derived materials formed a continuum from well-preserved to degraded states.
“That was genuinely surprising,” Klenner said. “The method captured not only the distinction between life and nonlife, but also degrees of preservation and alteration.”
Even heavily degraded biological samples retained traces of that organization. Fossilized dinosaur eggshells analyzed in the study, for example, still carried detectable statistical signatures shaped by ancient life.
The researchers emphasize that no single method is likely to prove the existence of extraterrestrial life on its own.
“Any future claim of having found life would require multiple independent lines of evidence, interpreted within the geological and chemical context of a planetary environment,” Klenner said.
Still, the team believes their framework could become an important new tool for future missions.
“Our approach is one more way to assess whether life may have been there,” Klenner said. “And if different techniques all point in the same direction, then that becomes very powerful.”
Astrobiology, Genomics, Biochemistry,
Stay Informed With the Latest & Most Important News
Previous Post
Next Post
-
 01Two Black Holes Observed Circling Each Other for the First Time
01Two Black Holes Observed Circling Each Other for the First Time -
 02From Polymerization-Enabled Folding and Assembly to Chemical Evolution: Key Processes for Emergence of Functional Polymers in the Origin of Life
02From Polymerization-Enabled Folding and Assembly to Chemical Evolution: Key Processes for Emergence of Functional Polymers in the Origin of Life -
 03Astronomy 101: From the Sun and Moon to Wormholes and Warp Drive, Key Theories, Discoveries, and Facts about the Universe (The Adams 101 Series)
03Astronomy 101: From the Sun and Moon to Wormholes and Warp Drive, Key Theories, Discoveries, and Facts about the Universe (The Adams 101 Series) -
 04True Anomaly hires former York Space executive as chief operating officer
04True Anomaly hires former York Space executive as chief operating officer -
 05Φsat-2 begins science phase for AI Earth images
05Φsat-2 begins science phase for AI Earth images -
 06Hurricane forecasters are losing 3 key satellites ahead of peak storm season − a meteorologist explains why it matters
06Hurricane forecasters are losing 3 key satellites ahead of peak storm season − a meteorologist explains why it matters -
 07Binary star systems are complex astronomical objects − a new AI approach could pin down their properties quickly
07Binary star systems are complex astronomical objects − a new AI approach could pin down their properties quickly