Now Reading: Identifying Amino Acid Isomers With Mass Spectrometry On Icy Ocean
-
01
Identifying Amino Acid Isomers With Mass Spectrometry On Icy Ocean
Identifying Amino Acid Isomers With Mass Spectrometry On Icy Ocean
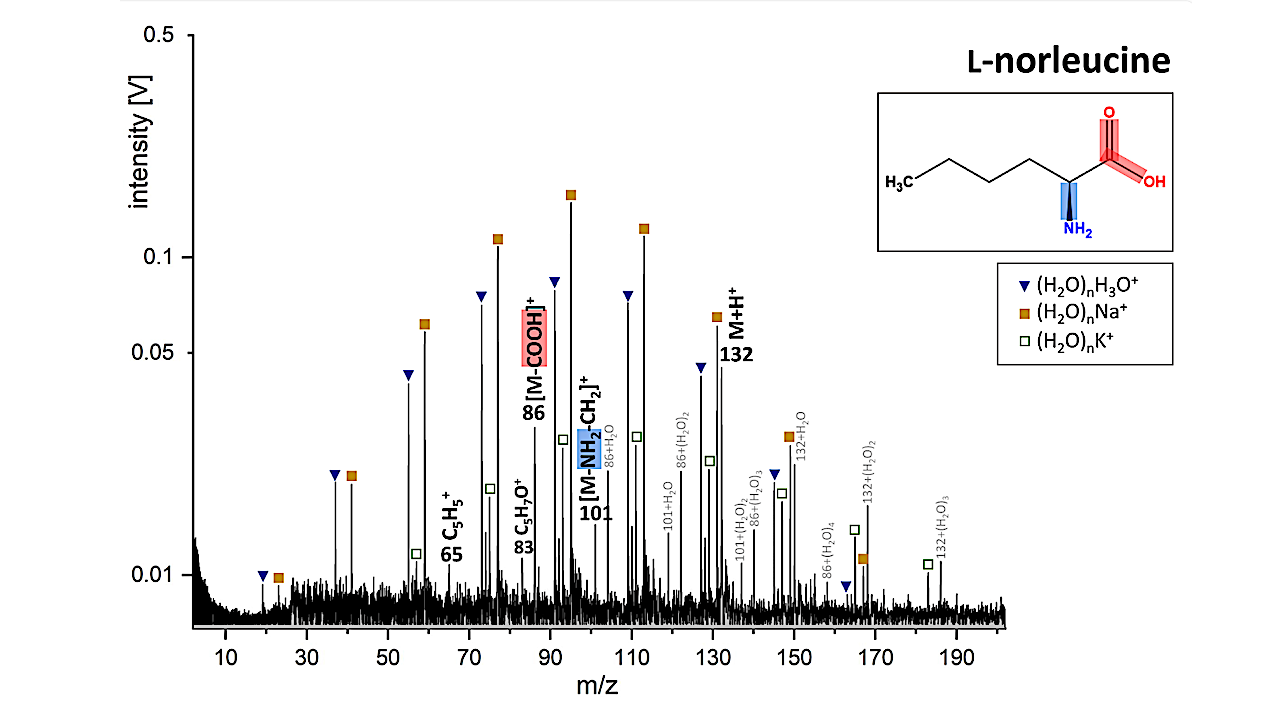
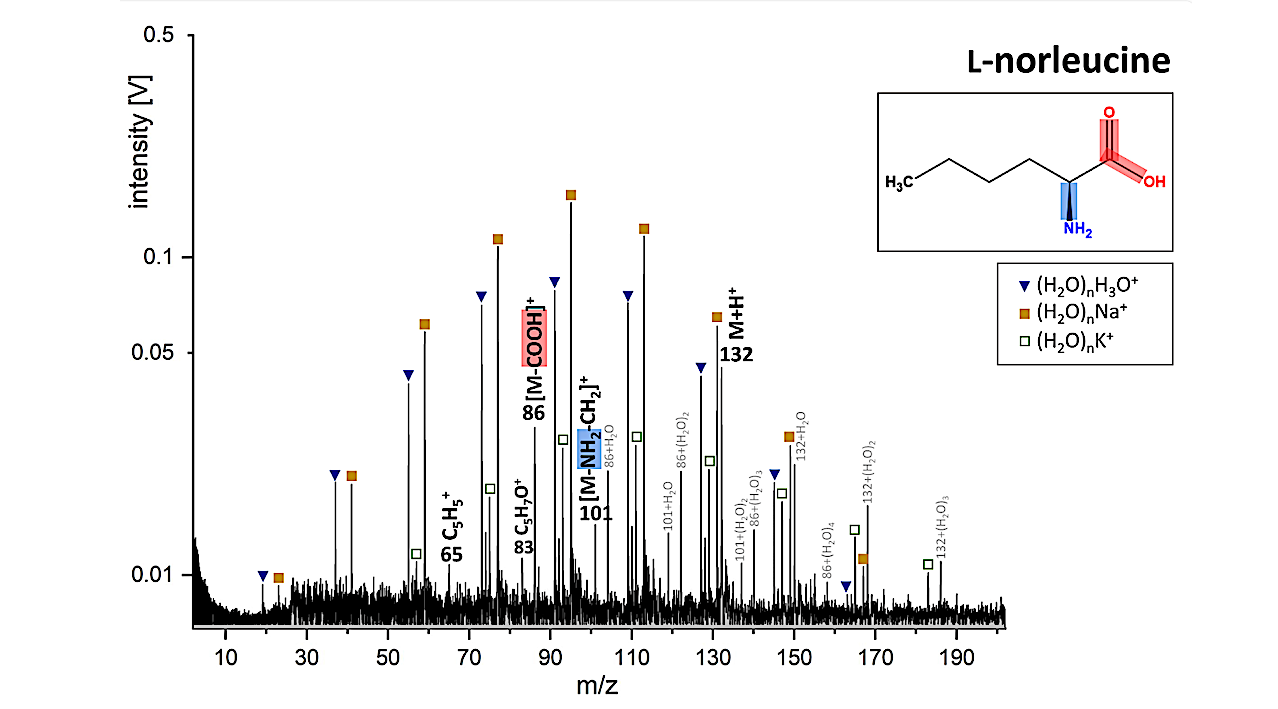
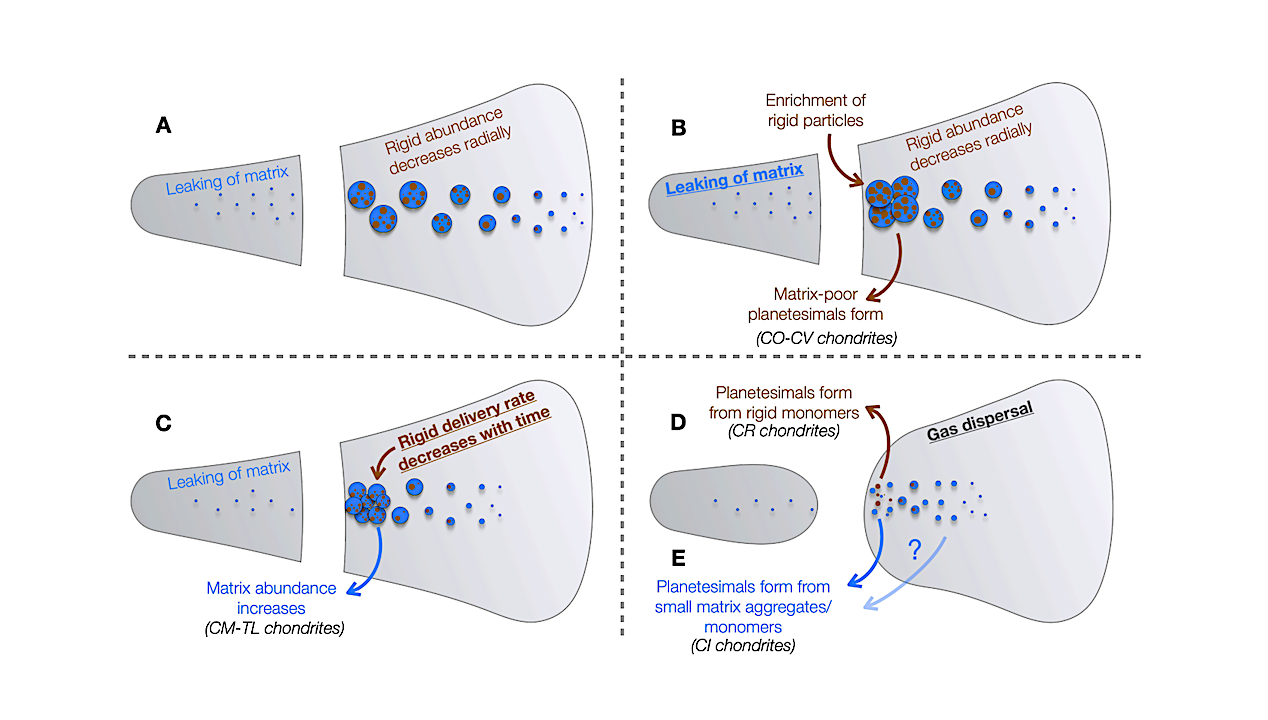
LILBID cationic mass spectrum of l-norleucine simulating an impact velocity of 5–7 km/s. The protonated molecular peak of l-norleucine is labeled “M+H+.” Fragment ions are identified in the mass spectrum and represented in the molecular structure as well. Ions from the matrix (pure water clusters and sodium and potassium water clusters) are labeled with blue triangles and yellow and green squares. — Astrobiology via x-mol.net
Enceladus and Europa are compelling targets for astrobiology investigations due to their potentially habitable subsurface oceans connected to the icy surface by geological processes.
Both moons emit ice grains either ejected via micrometeoroid surface impacts or erupted from their interiors through plume activity. These grains can be sampled by spacecraft flybys, and their composition can be analyzed by impact ionization mass spectrometers, such as the SUrface Dust Analyzer (SUDA) onboard Europa Clipper, or similar instruments proposed for future Enceladus missions. These instruments can identify potential biomolecules, such as amino acids, down to nanomolar concentrations, as demonstrated through previous laboratory experiments.
However, the identical masses of isomeric compounds could hinder the mass spectrometric identification and assignment of molecular biosignatures. Here, we investigate the general capability of impact ionization mass spectrometry to distinguish between isomeric compounds, validated with a test case of eight amino acid isomers with an identical molecular mass of 131.173 u and formula C6H13NO2, using quantum chemical calculations.
We show that the amino acid isomers (including diastereoisomers) can be uniquely identified due to their distinct mass spectral features and fragmentation patterns, explained through intramolecular hydrogen bonding and other structural specificities of the individual isomers.
Importantly, α-amino acids can be clearly differentiated from non-α-amino acids owing to several major mass spectral features. We show that SUDA-type instruments have sufficient capabilities to differentiate certain isomers and to identify biosignatures from ocean worlds with high confidence.
Astrobiology,
Stay Informed With the Latest & Most Important News
-
 01Two Black Holes Observed Circling Each Other for the First Time
01Two Black Holes Observed Circling Each Other for the First Time -
 02From Polymerization-Enabled Folding and Assembly to Chemical Evolution: Key Processes for Emergence of Functional Polymers in the Origin of Life
02From Polymerization-Enabled Folding and Assembly to Chemical Evolution: Key Processes for Emergence of Functional Polymers in the Origin of Life -
 03Astronomy 101: From the Sun and Moon to Wormholes and Warp Drive, Key Theories, Discoveries, and Facts about the Universe (The Adams 101 Series)
03Astronomy 101: From the Sun and Moon to Wormholes and Warp Drive, Key Theories, Discoveries, and Facts about the Universe (The Adams 101 Series) -
 04True Anomaly hires former York Space executive as chief operating officer
04True Anomaly hires former York Space executive as chief operating officer -
 05Φsat-2 begins science phase for AI Earth images
05Φsat-2 begins science phase for AI Earth images -
 06Hurricane forecasters are losing 3 key satellites ahead of peak storm season − a meteorologist explains why it matters
06Hurricane forecasters are losing 3 key satellites ahead of peak storm season − a meteorologist explains why it matters -
 07Binary star systems are complex astronomical objects − a new AI approach could pin down their properties quickly
07Binary star systems are complex astronomical objects − a new AI approach could pin down their properties quickly